
上海交通大学学报(医学版) ›› 2026, Vol. 46 ›› Issue (3): 275-290.doi: 10.3969/j.issn.1674-8115.2026.03.002
王俊骁1, 王雨泽2, 张沙沙2, 袁雨2, 张冰2, 李若谷1( )
)
收稿日期:2025-10-13
接受日期:2025-12-19
出版日期:2026-03-28
发布日期:2026-03-30
通讯作者:
李若谷,主任医师,博士;电子信箱:13564565961@163.com。基金资助:
Wang Junxiao1, Wang Yuze2, Zhang Shasha2, Yuan Yu2, Zhang Bing2, Li Ruogu1( )
)
Received:2025-10-13
Accepted:2025-12-19
Online:2026-03-28
Published:2026-03-30
Contact:
Li Ruogu, E-mail: 13564565961@163.com.Supported by:摘要:
目的·利用单细胞核RNA测序(single-nucleus RNA sequencing,snRNA-seq)技术,探究核纤层蛋白A/C基因第195位天冬酰胺替换为赖氨酸(Lamin A/C p.Asn195Lys,Lmna N195K)突变导致扩张型心肌病(dilated cardiomyopathy,DCM)小鼠模型心脏细胞的组成和分子特征变化,明确该突变对心脏多细胞生态系统的影响,并从细胞类型比例、基因表达谱、细胞间通信等方面系统描述其病理表型。方法·通过成簇规律间隔短回文重复序列及其相关蛋白9(clustered regularly interspaced short palindromic repeats-associated protein 9,CRISPR-Cas9)技术构建Lmna N195K纯合突变小鼠模型,并设立野生型对照。采用心脏超声和心电图评估心功能参数,通过苏木精-伊红(hematoxylin and eosin,HE)染色、天狼星红染色、小麦胚芽凝集素(wheat germ agglutinin,WGA)染色和末端脱氧核苷酸转移酶介导的dUTP缺口末端标记(terminal deoxynucleotidyl transferase dutp nick end labeling,TUNEL)凋亡检测分析心脏组织的病理变化。提取心脏组织细胞核进行snRNA-seq,使用Seurat(v4.4.0)进行细胞聚类、差异表达分析和细胞亚群鉴定,拟时序分析追踪细胞状态转变,CellChat(v1.6.1)推断细胞间通信网络。实时荧光定量逆转录聚合酶链反应(quantitative reverse transcription polymerase chain reaction,RT-qPCR)及蛋白质印迹法(Western blotting,WB)分别检测纤维化标志物和LMNA蛋白表达水平。结果·Lmna N195K纯合突变小鼠表现出心腔扩张、收缩功能下降、严重心脏纤维化和高死亡率。组织学分析显示心肌细胞排列紊乱、纤维化面积显著增加、凋亡细胞增多,且突变组中LMNA蛋白表达降低。snRNA-seq共鉴定出9种心脏细胞类型,包括心肌细胞、内皮细胞、成纤维细胞等。突变组中代谢型心肌细胞(CM2)比例显著降低(P=0.005),收缩型心肌细胞(CM3)比例上升(P=0.004);成纤维细胞几乎全部转化为促纤维化亚群FB4(P=0.001);内皮细胞中高细胞外基质(extracellular matrix,ECM)分泌亚群EC3比例增加(P=0.007),提示内皮-间质转化(endothelial-to-mesenchymal transition,EndoMT)发生。差异表达分析发现多个与纤维化、能量代谢和心脏损伤相关的基因表达异常。细胞间通信分析显示,突变组中骨膜蛋白(periostin,POSTN)-整合素(integrin)、血管内皮生长因子A(vascular endothelial growth factor A,VEGFA)-血管内皮生长因子受体1(vascular endothelial growth factor receptor 1,VEGFR1)以及层粘连蛋白亚基α2(laminin subunit alpha 2,LAMA2)-整合素(integrin)等受体-配体通路显著激活。结论·Lmna N195K突变引起心脏多细胞生态系统紊乱,主要表现为心肌细胞亚群比例失衡(代谢型减少、收缩型增加)、成纤维细胞向促纤维化表型转化、内皮细胞发生EndoMT并增强ECM分泌能力。这些变化通过异常的细胞间通信网络进一步协同促进心脏纤维化和病理性重塑,最终导致DCM的发生与发展。该研究从单细胞水平系统提示了Lmna N195K突变对心脏细胞组成和功能的影响,为理解该突变致DCM的细胞基础提供了重要数据支持。
中图分类号:
王俊骁, 王雨泽, 张沙沙, 袁雨, 张冰, 李若谷. 基于单细胞核RNA测序的Lmna N195K突变心肌病小鼠心脏特征分析[J]. 上海交通大学学报(医学版), 2026, 46(3): 275-290.
Wang Junxiao, Wang Yuze, Zhang Shasha, Yuan Yu, Zhang Bing, Li Ruogu. Analysis of cardiac features in Lmna N195K mutant mice using single-nucleus RNA sequencing[J]. Journal of Shanghai Jiao Tong University (Medical Science), 2026, 46(3): 275-290.
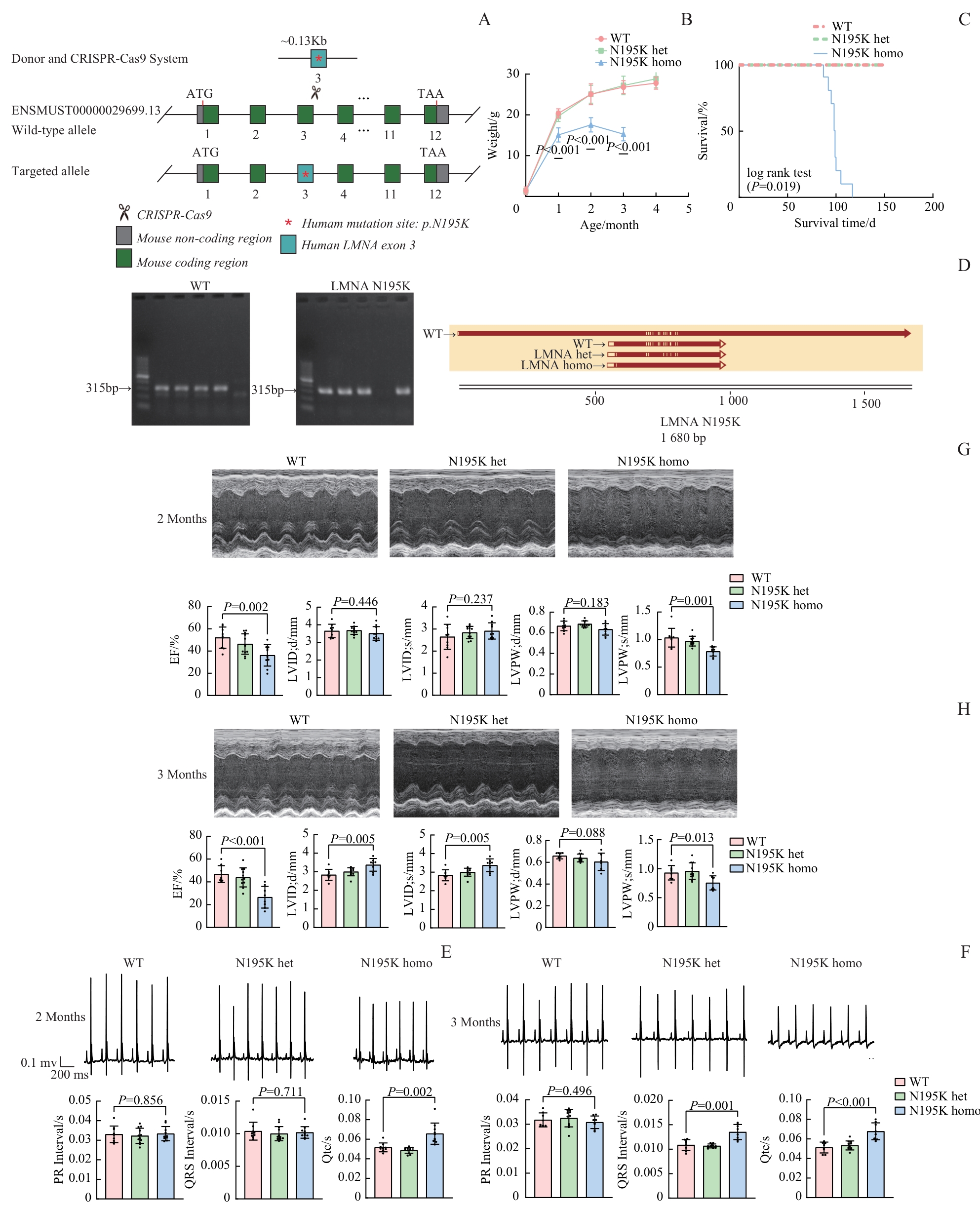
图1 LMNA p.N195K 基因敲入小鼠的构建及心脏功能表征Note: A. Schematic illustration of the CRISPR-Cas9 strategy utilized to generate Lmna p.N195K knock-in mice. The wild-type allele (ENSMUST0000003689.13) and the targeted allele containing the human p.N195K mutation in exon 3 are shown. B. Body weight curves of WT, N195K heterozygous (het), and homozygous (homo) mice at the age of 3‒4 months. C. Kaplan-Meier survival curves of WT, N195K het, and homo mice (log-rank test, P<0.001). D. Genotyping of Lmna N195K mutant mice (left: WT; right: Lmna N195K mutant); sequence validation of the corresponding alleles is shown below. E/F. Representative electrocardiograms and quantitative parameters (PR, QRS duration, QTc) in WT, N195K het, and homo mice at 2 (E) and 3 (F) months of age (x±s). G/H. Representative echocardiographic images and cardiac parameters, including EF, LVID; d, LVID; s, LVPW; d, and LVPW; s, in WT, N195K het, and homo mice at 2 (G) and 3 (H) months of age.
Fig 1 Generation of LMNA p.N195K knock-in mice and characterization of cardiac function
| Gene | Forward primer sequence (5′→3′) | Reverse primer sequence (5′→3′) |
|---|---|---|
| Anp | AGCTGGATCTTCGTAGGCTC | GCCGGTAGAAGATGAGGTCA |
| Bnp | GAGGTCACTCCTATCCTCTGG | GCCATTTCCTCCGACTTTTCTC |
| Myh6 | CAAGCTCACTTGAAGGACACC | CACGATGGCGATGTTCTC |
| Myh7 | AACCAGACGGCACTGAAGAG | TGCCCACTTTGACTCTAGGATG |
| Col1a1 | GCTCCTCTTAGGGGCCACT | CCACGTCTCACCATTGGGG |
| Col1a2 | GTAACTTCGTGCCTAGCAACA | CCTTTGTCAGAATACTGAGCAGC |
| Col2a1 | GGGAATGTCCTCTGCGATGAC | GAAGGGGATCTCGGGGTTG |
| Col4a1 | CTGGCACAAAAGGGACGAG | ACGTGGCCGAGAATTTCACC |
| Col4a2 | GACCGAGTGCGGTTCAAAG | CGCAGGGCACATCCAACTT |
| Tgfb1 | CTCCCGTGGCTTCTAGTGC | GCCTTAGTTTGGACAGGATCTG |
| Tgfb2 | TCGACATGGATCAGTTTATGCG | CCCTGGTACTGTTGTAGATGGA |
| Tgfb3 | CAGGCCAGGGTAGTCAGAG | ATTTCCAGCCTAGATCCTGCC |
表1 RT-qPCR引物序列
Tab 1 Primer sequences for RT-qPCR
| Gene | Forward primer sequence (5′→3′) | Reverse primer sequence (5′→3′) |
|---|---|---|
| Anp | AGCTGGATCTTCGTAGGCTC | GCCGGTAGAAGATGAGGTCA |
| Bnp | GAGGTCACTCCTATCCTCTGG | GCCATTTCCTCCGACTTTTCTC |
| Myh6 | CAAGCTCACTTGAAGGACACC | CACGATGGCGATGTTCTC |
| Myh7 | AACCAGACGGCACTGAAGAG | TGCCCACTTTGACTCTAGGATG |
| Col1a1 | GCTCCTCTTAGGGGCCACT | CCACGTCTCACCATTGGGG |
| Col1a2 | GTAACTTCGTGCCTAGCAACA | CCTTTGTCAGAATACTGAGCAGC |
| Col2a1 | GGGAATGTCCTCTGCGATGAC | GAAGGGGATCTCGGGGTTG |
| Col4a1 | CTGGCACAAAAGGGACGAG | ACGTGGCCGAGAATTTCACC |
| Col4a2 | GACCGAGTGCGGTTCAAAG | CGCAGGGCACATCCAACTT |
| Tgfb1 | CTCCCGTGGCTTCTAGTGC | GCCTTAGTTTGGACAGGATCTG |
| Tgfb2 | TCGACATGGATCAGTTTATGCG | CCCTGGTACTGTTGTAGATGGA |
| Tgfb3 | CAGGCCAGGGTAGTCAGAG | ATTTCCAGCCTAGATCCTGCC |
| Primer name | Primer sequence |
|---|---|
| ms-lmna195-f(WT) | CTGAGAACAGGCTACAGACGC |
| h-lmna195-f(Lmna N195K) | GAGAACAGGCTGCAGACCA |
| lmna195-r(WT/ Lmna N195K) | GCTCTCAAACTCTCGCTGCT |
| LMNA195-tF1(Sanger) | GTGCCATGGGTAGAATGTGGGT |
| LMNA195-tR1(Sanger) | GAAAGGGTTAAGGGGAGACACT |
表2 基因型鉴定所用引物序列
Tab 2 Primer sequences used for genotyping
| Primer name | Primer sequence |
|---|---|
| ms-lmna195-f(WT) | CTGAGAACAGGCTACAGACGC |
| h-lmna195-f(Lmna N195K) | GAGAACAGGCTGCAGACCA |
| lmna195-r(WT/ Lmna N195K) | GCTCTCAAACTCTCGCTGCT |
| LMNA195-tF1(Sanger) | GTGCCATGGGTAGAATGTGGGT |
| LMNA195-tR1(Sanger) | GAAAGGGTTAAGGGGAGACACT |
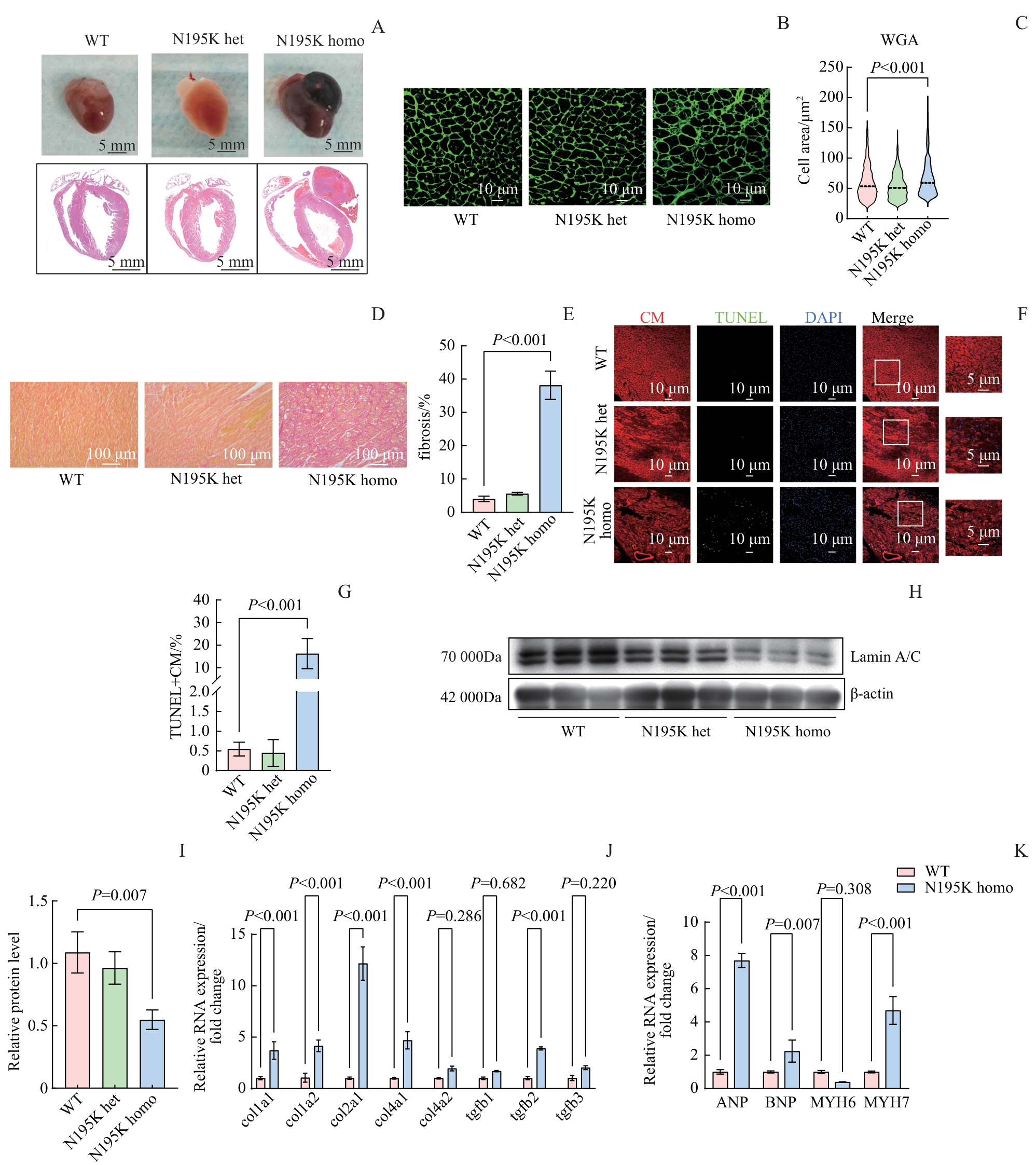
图2 Lmna N195K 小鼠心脏组织病理学及分子改变Note: A. Gross morphology and HE staining of cardiac tissues (Upper panel: gross morphology; lower panel: HE staining. Scale bar: 5 mm for gross morphology, 1 mm for HE staining). B. WGA staining of cardiomyocyte membranes. Scale bar: 10 μm. C. Quantification of cardiomyocyte cross-sectional area from panel (B) (n≥600 cells). D. Sirius red staining for assessment of cardiac fibrosis. Scale bar: 100 μm. E. Quantification of fibrotic area from panel (D). F. TUNEL/DAPI/α-sarcomeric actin co-staining to identify apoptotic cardiomyocytes. Scale bar: 10 μm (20×), 5 μm (60×). G. Apoptotic cardiomyocytes (TUNEL+ CM) from panel (F). H. Immunoblot analysis of Lamin A/C protein in cardiac tissue; β-actin served as the loading control. I. Densitometric quantification of Lamin A/C protein relative to β-actin. J. Expression of fibrosis-related genes (Col1a1, Col1a2, Col2a1, Col4a1, Col4a2, Tgfb1, Tgfb2, Tgfb3) evaluated by quantitative PCR. K. Expression of heart failure markers (Anp, Bnp, Myh6, Myh7) assessed by quantitative PCR. Col1a1—Collagen, type I, alpha 1; Tgfb1-Transforming growth factor, beta 1.
Fig 2 Histopathological and molecular alterations in the hearts of Lmna N195K mice
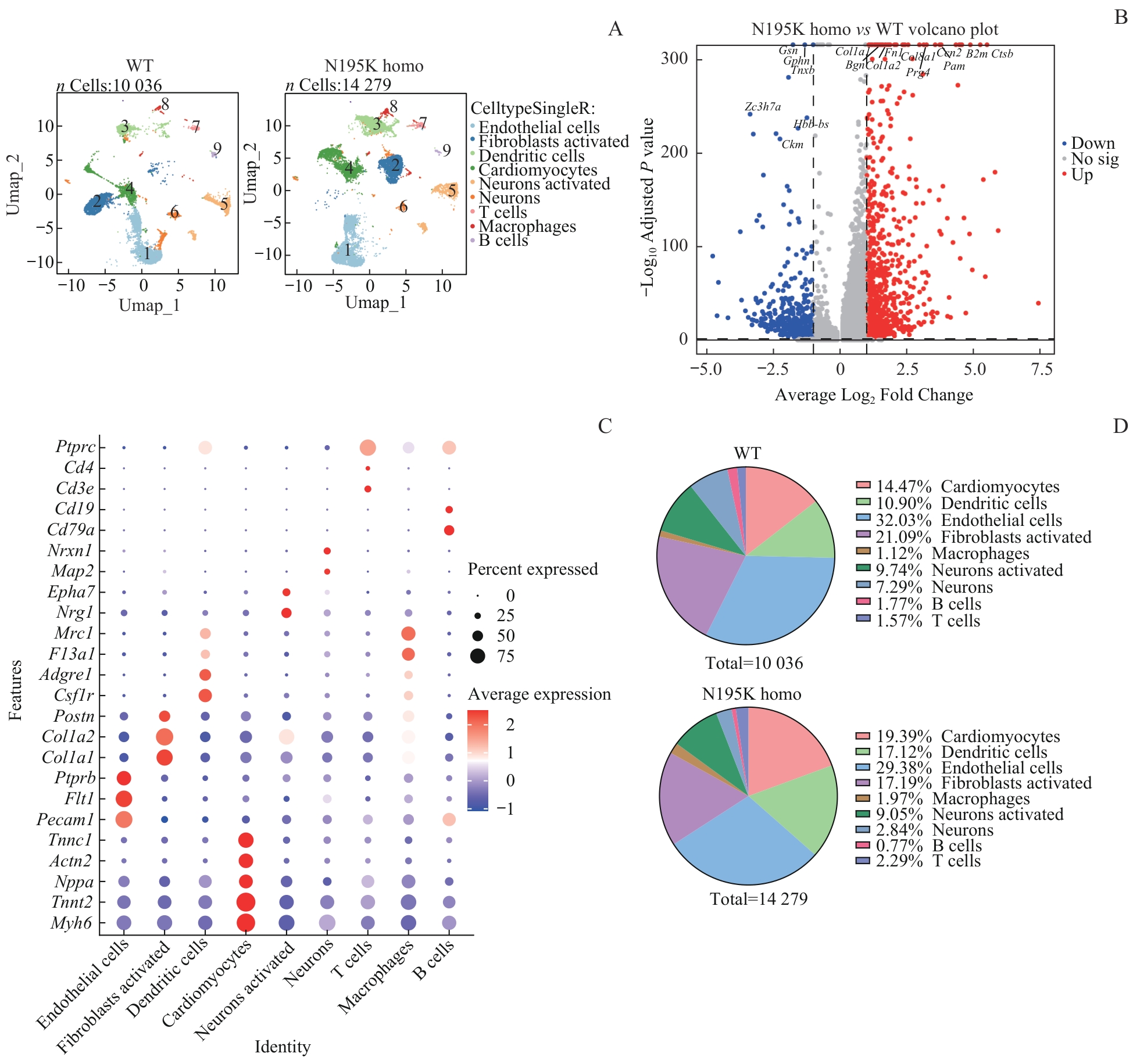
图3 单细胞核RNA测序总体分析Note: A. UMAP plots of annotated cardiac cell nuclei from WT and Lmna N195K homozygous mice (cell clusters: 1-cardiomyocytes; 2-endothelial cells; 3-fibroblasts; 4-dendritic cells; 5-macrophages; 6-neurons activated; 7-neurons; 8-B cells; 9-T cells; WT: 10 036 nuclei; Lmna N195K homo: 14 279 nuclei). B. Volcano plot of differentially expressed genes (DEGs) between Lmna N195K homozygous and WT groups (red dots: upregulated DEGs; blue dots: downregulated DEGs). C. Bubble plot of DEGs across distinct cell types. D. Pie charts showing changes in the proportions of different cell types between WT and Lmna N195K homo groups (WT: 10 036 nuclei; Lmna N195K homo: 14 279 nuclei).
Fig 3 Global analysis of single-nucleus RNA sequencing (snRNA-seq)
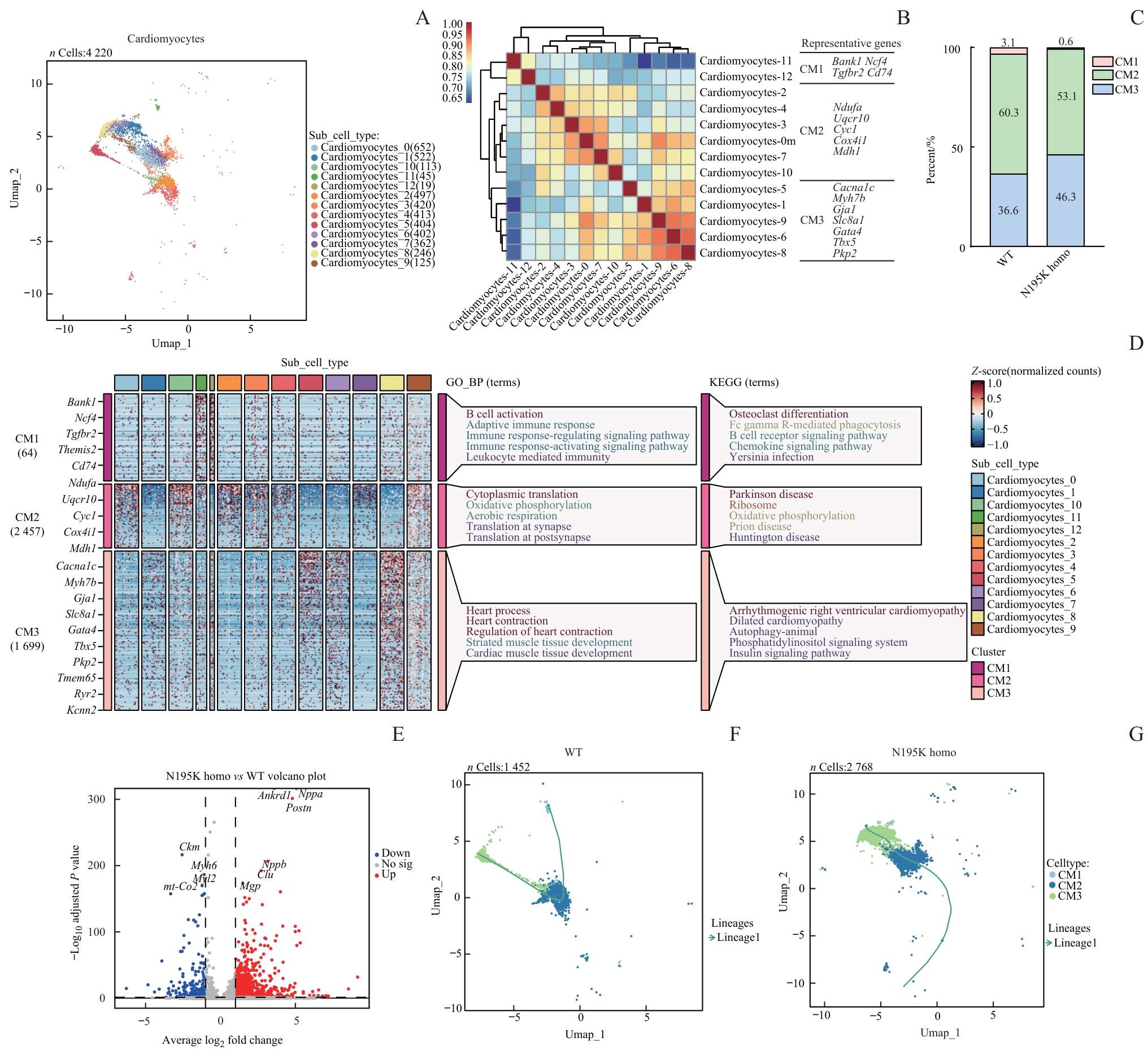
图4 单核RNA测序提示 Lmna N195K 纯合突变导致的心肌细胞表达重塑与状态转变Note: A. UMAP plots of annotated cardiomyocyte nuclei (nuclei: 22 020) from WT and Lmna N195K homozygous mice. B. Spearman correlation analysis of all cardiomyocytes, showing expression correlations and transcriptional similarities among different subpopulations. C. Thirteen cardiomyocyte subclusters integrated (or categorized) into three major groups (CM1, CM2, and CM3), with proportions compared between WT and N195K homozygous groups, demonstrating LMNA mutation-induced shifts in cardiomyocyte composition. D. Bubble plot and GO enrichment analysis identifying significantly enriched biological processes and pathways in cardiomyocytes from LMNA mutant mice. E. Volcano plot of differentially expressed genes in cardiomyocytes between N195K homozygous and WT mice (red dots: upregulated genes; blue dots: downregulated genes). F/G. Pseudotime trajectory analysis of cardiomyocytes in WT and N195K homozygous groups, depicting transcriptional evolution under pathological conditions and potential cell fate transitions.
Fig 4 Single-nucleus RNA sequencing reveals transcriptional remodeling and state transitions of cardiomyocytes induced by the Lmna N195K homozygous mutation
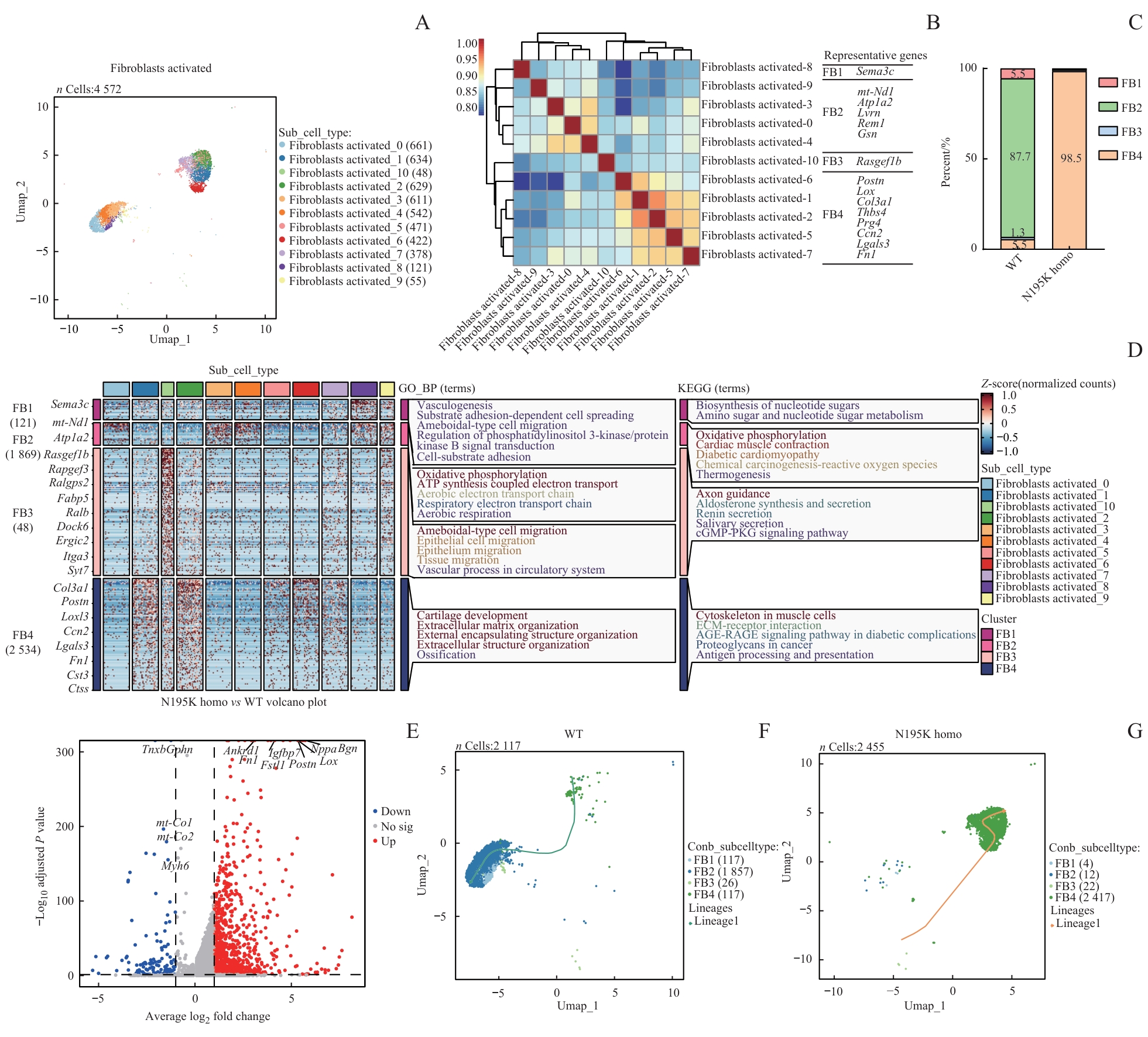
图5 基于单核RNA测序的 Lmna N195K 突变导致心脏成纤维细胞状态改变及功能重塑分析Note: A. UMAP plots of annotated cardiac fibroblast nuclei from WT and Lmna N195K homozygous mice. B. Spearman correlation analysis of all cardiac fibroblasts showing expression correlations and transcriptional similarities among different subpopulations. C. Eleven cardiac fibroblast subclusters integrated (or categorized) into four major groups (FB1, FB2, FB3, and FB4), demonstrating LMNA mutation-induced shifts in fibroblast composition. D. GO enrichment analysis identifying significantly enriched biological processes and pathways in cardiac fibroblasts from LMNA mutant mice. E. Volcano plot of differentially expressed genes in cardiac fibroblasts between N195K homozygous and WT mice. F/G. Pseudotime trajectory analysis of cardiac fibroblasts in WT and N195K homozygous groups, depicting transcriptional evolution under pathological conditions and potential cell fate transitions.
Fig 5 Analysis of state alteration and functional remodeling of cardiac fibroblasts induced by Lmna N195K mutation based on snRNA-seq
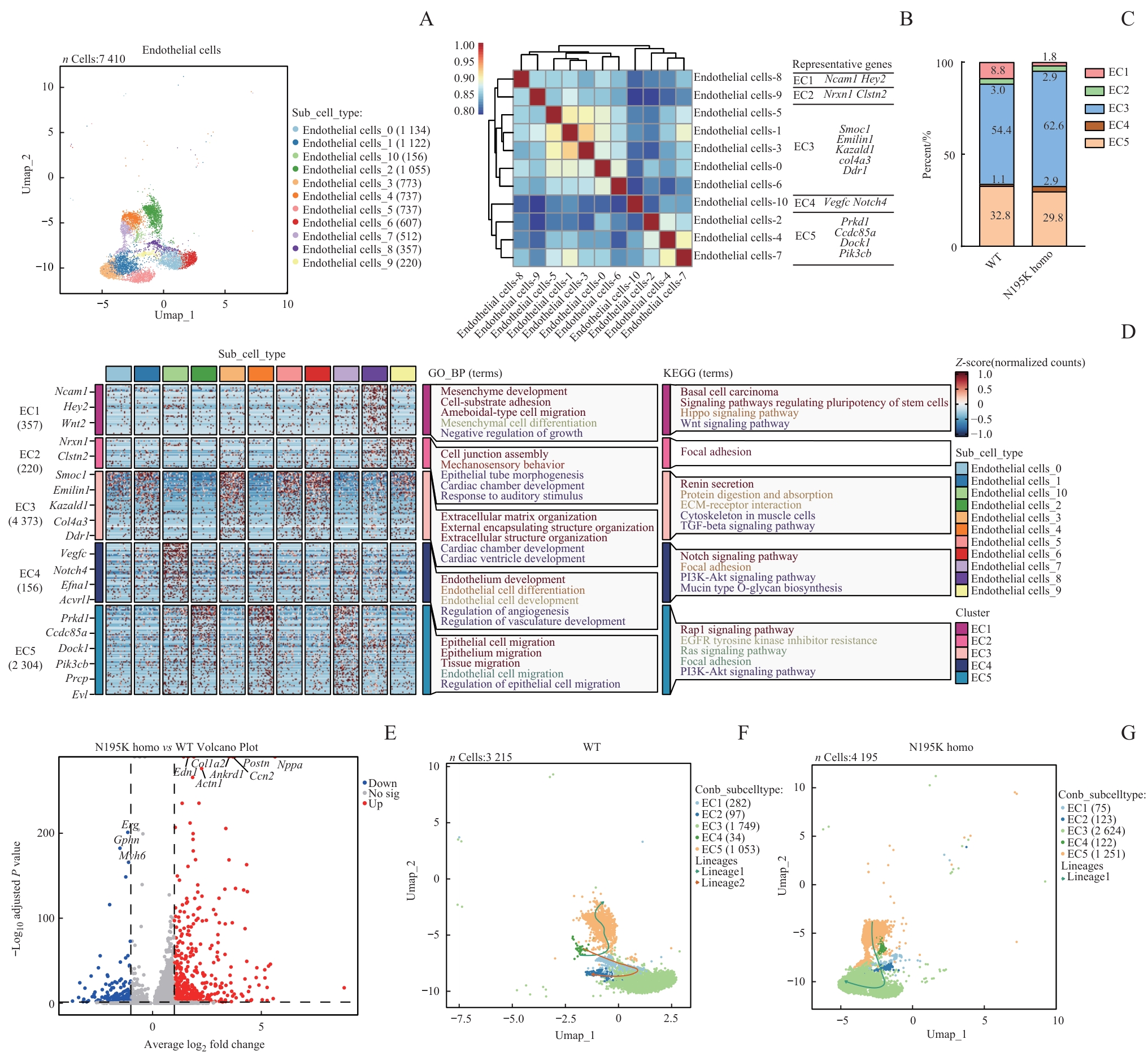
图6 基于单核RNA测序的 Lmna N195K 突变致心脏内皮细胞状态改变及功能重塑分析Note: A. UMAP plots of annotated cardiac endothelial cell nuclei from WT and Lmna N195K homozygous mice. B. Spearman correlation analysis of all cardiac endothelial cells, showing expression correlations and transcriptional similarities among different subpopulations. C. Integration of 11 cardiac endothelial cell subclusters into five major groups (EC1, EC2, EC3, EC4, EC5) demonstrating LMNA mutation-induced alterations in endothelial cell composition. D. GO enrichment analysis identifying significantly enriched biological processes and pathways in cardiac endothelial cells from LMNA mutant mice. E. Volcano plot of differentially expressed genes in cardiac endothelial cells between N195K homozygous and WT mice. F/G. Pseudotime trajectory analysis of cardiac endothelial cells in WT and N195K homozygous groups, illustrating transcriptional evolution under pathological conditions and potential cell fate transitions.
Fig 6 Analysis of state alteration and functional remodeling of cardiac endothelial cells induced by Lmna N195K mutation based on snRNA-seq
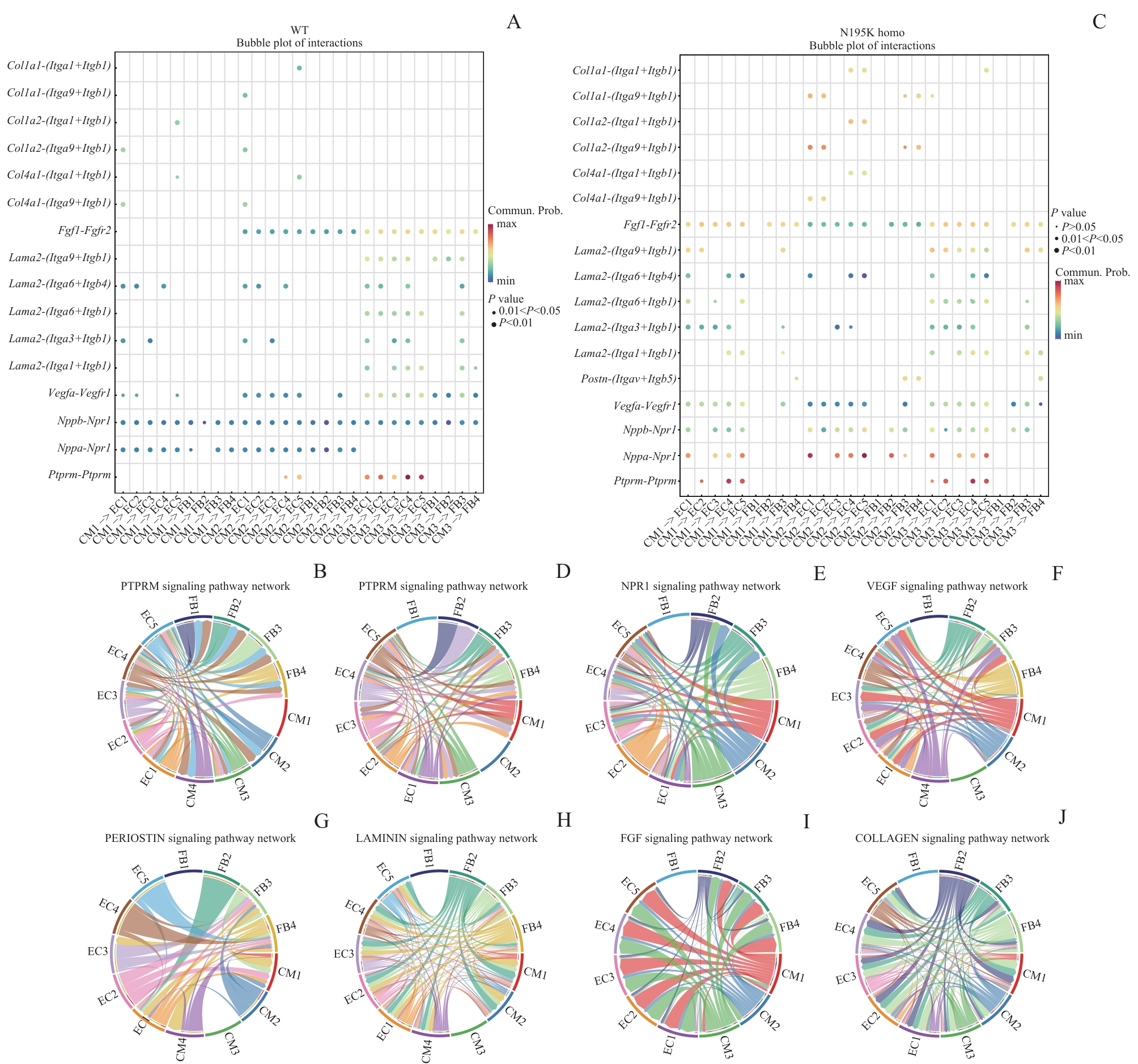
图7 WT和 Lmna N195K 纯合突变型小鼠心脏信号通路的受体-配体联系网络Note: A. Bubble plot of receptor-ligand interactions in cardiac tissue in the WT group. B. Circos plot of PTPRM-related receptor-ligand interactions in cardiac tissue in the WT group. C. Bubble plot of receptor-ligand interactions in cardiac tissue in the Lmna N195K homozygous group. D—J. Circos plots of receptor-ligand interactions that were highly expressed in the Lmna N195K homozygous group.
Fig7 Receptor-ligand interaction networks in cardiac signaling pathways of WT and Lmna N195K homozygous mice
| [1] | Broers J L, Ramaekers F C, Bonne G, et al. Nuclear lamins: laminopathies and their role in premature ageing[J]. Physiol Rev, 2006, 86(3): 967-1008. |
| [2] | Fatkin D, MacRae C, Sasaki T, et al. Missense mutations in the rod domain of the lamin A/C gene as causes of dilated cardiomyopathy and conduction-system disease[J]. N Engl J Med, 1999, 341(23): 1715-1724. |
| [3] | Worman H J, Fong L G, Muchir A, et al. Laminopathies and the long strange trip from basic cell biology to therapy[J]. J Clin Invest, 2009, 119(7): 1825-1836. |
| [4] | van Berlo J H, de Voogt W G, van der Kooi A J, et al. Meta-analysis of clinical characteristics of 299 carriers of LMNA gene mutations: do lamin A/C mutations portend a high risk of sudden death [J]. J Mol Med (Berl), 2005, 83(1): 79-83. |
| [5] | Hasselberg N E, Haland T F, Saberniak J, et al. Lamin A/C cardiomyopathy: young onset, high penetrance, and frequent need for heart transplantation[J]. Eur Heart J, 2018, 39(10): 853-860. |
| [6] | Nishiuchi S, Makiyama T, Aiba T, et al. Gene-based risk stratification for cardiac disorders in LMNA mutation carriers[J]. Circ Cardiovasc Genet, 2017, 10(6): e001603. |
| [7] | Rosario K F, Karra R, Amos K, et al. LMNA cardiomyopathy: important considerations for the heart failure clinician[J]. J Card Fail, 2023, 29(12): 1657-1666. |
| [8] | Bank E M, Ben-Harush K, Feinstein N, et al. Structural and physiological phenotypes of disease-linked lamin mutations in C. elegans[J]. J Struct Biol, 2012, 177(1): 106-112. |
| [9] | Ito M, Katoh M, Sassa T, et al. LMNA Q353R mutation causes dilated cardiomyopathy through impaired vitamin D signaling[J]. Circulation, 2024, 150(12): 971-974. |
| [10] | Lee J, Termglinchan V, Diecke S, et al. Activation of PDGF pathway links LMNA mutation to dilated cardiomyopathy[J]. Nature, 2019, 572(7769): 335-340. |
| [11] | Mounkes L C, Kozlov S V, Rottman J N, et al. Expression of an LMNA-N195K variant of A-type lamins results in cardiac conduction defects and death in mice[J]. Hum Mol Genet, 2005, 14(15): 2167-2180. |
| [12] | Markandeya Y S, Tsubouchi T, Hacker T A, et al. Inhibition of late sodium current attenuates ionic arrhythmia mechanism in ventricular myocytes expressing LaminA-N195K mutation[J]. Heart Rhythm, 2016, 13(11): 2228-2236. |
| [13] | Laurini E, Martinelli V, Lanzicher T, et al. Biomechanical defects and rescue of cardiomyocytes expressing pathologic nuclear lamins[J]. Cardiovasc Res, 2018, 114(6): 846-857. |
| [14] | Neubauer S. The failing heart: an engine out of fuel[J]. N Engl J Med, 2007, 356(11): 1140-1151. |
| [15] | Lazaropoulos M P, Elrod J W. Mitochondria in pathological cardiac remodeling[J]. Curr Opin Physiol, 2022, 25: 100489. |
| [16] | Doenst T, Nguyen T D, Abel E D. Cardiac metabolism in heart failure: implications beyond ATP production[J]. Circ Res, 2013, 113(6): 709-724. |
| [17] | Scuruchi M, Mannino F, Imbesi C, et al. Biglycan involvement in heart fibrosis: modulation of adenosine 2A receptor improves damage in immortalized cardiac fibroblasts[J]. Int J Mol Sci, 2023, 24(2): 1784. |
| [18] | Wei K, Serpooshan V, Hurtado C, et al. Epicardial FSTL1 reconstitution regenerates the adult mammalian heart[J]. Nature, 2015, 525(7570): 479-485. |
| [19] | Peters M C, di Martino S, Boelens T, et al. Follistatin-like 1 promotes proliferation of matured human hypoxic iPSC-cardiomyocytes and is secreted by cardiac fibroblasts[J]. Mol Ther Methods Clin Dev, 2022, 25: 3-16. |
| [20] | Koenig A L, Shchukina I, Amrute J, et al. Single-cell transcriptomics reveals cell-type-specific diversification in human heart failure[J]. Nat Cardiovasc Res, 2022, 1(3): 263-280. |
| [21] | Qian C, Dong G L, Yang C M, et al. Broadening horizons: molecular mechanisms and disease implications of endothelial-to-mesenchymal transition[J]. Cell Commun Signal, 2025, 23(1): 16. |
| [22] | Kovacic J C, Dimmeler S, Harvey R P, et al. Endothelial to mesenchymal transition in cardiovascular disease: JACC state-of-the-art review[J]. J Am Coll Cardiol, 2019, 73(2): 190-209. |
| [23] | Souders C A, Bowers S L, Baudino T A. Cardiac fibroblast: the renaissance cell[J]. Circ Res, 2009, 105(12): 1164-1176. |
| [24] | Kedzierski R M, Yanagisawa M. Endothelin system: the double-edged sword in health and disease[J]. Annu Rev Pharmacol Toxicol, 2001, 41: 851-876. |
| [25] | Oka T, Xu J, Kaiser R A, et al. Genetic manipulation of periostin expression reveals a role in cardiac hypertrophy and ventricular remodeling[J]. Circ Res, 2007, 101(3): 313-321. |
| [26] | Braile M, Marcella S, Cristinziano L, et al. VEGF-A in cardiomyocytes and heart diseases[J]. Int J Mol Sci, 2020, 21(15): 5294. |
| [27] | Shibuya M. Vascular endothelial growth factor (VEGF) and its receptor (VEGFR) signaling in angiogenesis: a crucial target for anti- and pro-angiogenic therapies[J]. Genes Cancer, 2011, 2(12): 1097-1105. |
| [28] | Aumailley M. The laminin family[J]. Cell Adh Migr, 2013, 7(1): 48-55. |
| [29] | Guo C, Willem M, Werner A, et al. Absence of α7 integrin in dystrophin-deficient mice causes a myopathy similar to Duchenne muscular dystrophy[J]. Hum Mol Genet, 2006, 15(6): 989-998. |
| [30] | Israeli-Rosenberg S, Manso A M, Okada H, et al. Integrins and integrin-associated proteins in the cardiac myocyte[J]. Circ Res, 2014, 114(3): 572-586. |
| [1] | 魏豪, 邱家俊, 颜景斌. 不同表达矩阵对筛选差异长链非编码RNA的影响[J]. 上海交通大学学报(医学版), 2022, 42(7): 911-918. |
| [2] | 邢冬梅, 朱明军, 吴泰相, 等. 干细胞移植治疗扩张型心肌病疗效和安全性的系统评价[J]. , 2012, 32(3): 321-. |
| [3] | 牛 琳, 陆颖静, 徐大地, 等. 儿童扩张型心肌病心腔血液超声背向散射参数的研究[J]. , 2009, 29(9): 1101-. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||