
Journal of Shanghai Jiao Tong University (Medical Science) ›› 2026, Vol. 46 ›› Issue (3): 358-367.doi: 10.3969/j.issn.1674-8115.2026.03.010
• Review • Previous Articles Next Articles
Tian Qijia, Jia Kangni, Yan Xiaoxiang( )
)
Received:2025-07-27
Accepted:2025-12-30
Online:2026-03-28
Published:2026-03-30
Contact:
Yan Xiaoxiang
E-mail:cardexyanxx@hotmail.com
Supported by:CLC Number:
Tian Qijia, Jia Kangni, Yan Xiaoxiang. Advances in CRISPR/Cas gene editing and its applications in cardiovascular diseases[J]. Journal of Shanghai Jiao Tong University (Medical Science), 2026, 46(3): 358-367.
Add to citation manager EndNote|Ris|BibTeX
URL: https://xuebao.shsmu.edu.cn/EN/10.3969/j.issn.1674-8115.2026.03.010
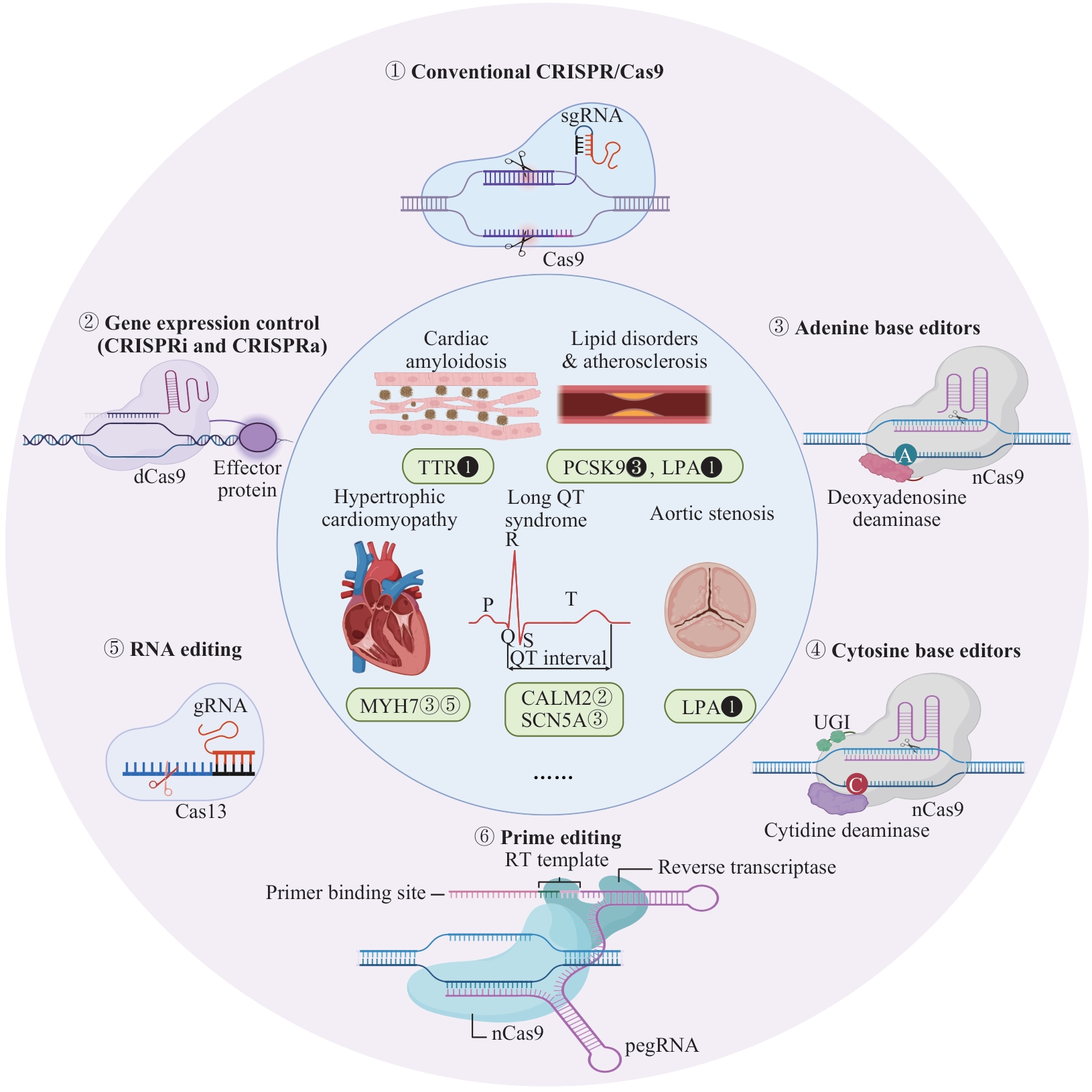
Fig 1 Schematic diagram of CRISPR/Cas applications in cardiovascular diseasesNote: Outer ring: 6 main categories of gene editing tools; inner ring: associated cardiovascular diseases and therapeutic targets. Markers adjacent to gene targets indicate the corresponding editing technologies (matching the outer ring numbering). Solid markers denote therapies in clinical trials; hollow markers indicate therapies supported only by in vivo animal evidence. CALM2—calmodulin 2; CRISPRi/a—CRISPR interference/activation; dCas9—dead Cas9; LPA—lipoprotein(a); MYH7—myosin heavy chain 7; nCas9—Cas9 nickase; PCSK9—proprotein convertase subtilisin/kexin type 9; pegRNA—prime editing guide RNA; RT—reverse transcription; SCN5A—sodium voltage-gated channel alpha subunit 5; TTR—transthyretin; UGI—uracil glycosylase inhibitor.
| Treatment | Gene editing method | Indication | Target gene | Delivery method | Clinical trial ID | Phase | Efficacy | Follow-up | Safety | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| NTLA-2001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06128629 | Ⅲ | TTR reduction: 89% (Day 28), 90% (Month 12) | 87% reduction sustained (Month 36) | Most common treatment-related AEs: mild infusion-related reactions (14%) and transient liver enzyme elevations (6%) | [ |
| YOLT-201 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06539208 | Ⅰ/Ⅱa | Trial ongoing | \ | \ | [ |
| ART001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | ChiCTR2400081216 | Ⅰ | Trial ongoing | \ | \ | [ |
| YOLT-101 | Base editing | Familial hypercholesterolemia | PCSK9 | GalNAc-LNP | NCT06458010 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 72.5% and LDL-C by 50.4% (Month 4) | Follow-up ongoing | Most common treatment-related AEs; mild infusion-related reactions (83%) and transient liver enzyme elevations (50%) | [ |
| VERVE-101 | Base editing | Familial hypercholesterolemia | PCSK9 | LNP | NCT05398029 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 47% and LDL-C by 55% (Day 28); sustained through 180 d | Follow-up ongoing | Treatment-related Grade 3 serious AEs: transient liver enzyme elevations and thrombocytopenia; mild infusion-related reactions were common; trial enrollment paused due to safety concerns | [ |
| VERVE-102 | Base editing | Familial hypercholesterolemia or premature coronary artery disease | PCSK9 | GalNAc-LNP | NCT06164730 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX320 | CRISPR/Cas9 knock out | Atherosclerosis or calcific aortic valve stenosis | LPA | LNP | ACTRN12623001095651 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX310 | CRISPR/Cas9 knock out | Refractory dyslipidemias | ANGPTL3 | LNP | ACTRN12623000809639 | Ⅰ | High-dose (0.8 mg/kg): ANGPTL3 reduced by 73.2%, LDL-C by 48.9%, and triglycerides by 55.2% (Day 60) | Follow-up ongoing | Most common treatment-related AEs: mild infusion-related reactions (20%) and transient liver enzyme elevations (7%) | [ |
| VERVE-201 | Base editing | Refractory hypercholesterolemia | ANGPTL3 | GalNAc-LNP | NCT06451770 | Ⅰ | Trial ongoing | \ | \ | [ |
Tab 1 Gene editing therapies for cardiovascular diseases in clinical trials
| Treatment | Gene editing method | Indication | Target gene | Delivery method | Clinical trial ID | Phase | Efficacy | Follow-up | Safety | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| NTLA-2001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06128629 | Ⅲ | TTR reduction: 89% (Day 28), 90% (Month 12) | 87% reduction sustained (Month 36) | Most common treatment-related AEs: mild infusion-related reactions (14%) and transient liver enzyme elevations (6%) | [ |
| YOLT-201 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06539208 | Ⅰ/Ⅱa | Trial ongoing | \ | \ | [ |
| ART001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | ChiCTR2400081216 | Ⅰ | Trial ongoing | \ | \ | [ |
| YOLT-101 | Base editing | Familial hypercholesterolemia | PCSK9 | GalNAc-LNP | NCT06458010 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 72.5% and LDL-C by 50.4% (Month 4) | Follow-up ongoing | Most common treatment-related AEs; mild infusion-related reactions (83%) and transient liver enzyme elevations (50%) | [ |
| VERVE-101 | Base editing | Familial hypercholesterolemia | PCSK9 | LNP | NCT05398029 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 47% and LDL-C by 55% (Day 28); sustained through 180 d | Follow-up ongoing | Treatment-related Grade 3 serious AEs: transient liver enzyme elevations and thrombocytopenia; mild infusion-related reactions were common; trial enrollment paused due to safety concerns | [ |
| VERVE-102 | Base editing | Familial hypercholesterolemia or premature coronary artery disease | PCSK9 | GalNAc-LNP | NCT06164730 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX320 | CRISPR/Cas9 knock out | Atherosclerosis or calcific aortic valve stenosis | LPA | LNP | ACTRN12623001095651 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX310 | CRISPR/Cas9 knock out | Refractory dyslipidemias | ANGPTL3 | LNP | ACTRN12623000809639 | Ⅰ | High-dose (0.8 mg/kg): ANGPTL3 reduced by 73.2%, LDL-C by 48.9%, and triglycerides by 55.2% (Day 60) | Follow-up ongoing | Most common treatment-related AEs: mild infusion-related reactions (20%) and transient liver enzyme elevations (7%) | [ |
| VERVE-201 | Base editing | Refractory hypercholesterolemia | ANGPTL3 | GalNAc-LNP | NCT06451770 | Ⅰ | Trial ongoing | \ | \ | [ |
| [1] | Mensah G A, Fuster V, Murray C J L, et al. Global burden of cardiovascular diseases and risks, 1990‒2022[J]. J Am Coll Cardiol, 2023, 82(25): 2350-2473. |
| [2] | Kim Y G, Cha J, Chandrasegaran S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain[J]. Proc Natl Acad Sci U S A, 1996, 93(3): 1156-1160. |
| [3] | Christian M, Cermak T, Doyle E L, et al. Targeting DNA double-strand breaks with TAL effector nucleases[J]. Genetics, 2010, 186(2): 757-761. |
| [4] | Jinek M, Chylinski K, Fonfara I, et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity[J]. Science, 2012, 337(6096): 816-821. |
| [5] | 中华医学会心血管病学分会心力衰竭学组, 中华心血管病杂志编辑委员会. 转甲状腺素蛋白心脏淀粉样变诊断与治疗中国专家共识[J]. 中华心血管病杂志, 2021, 49(4): 324-332. |
| Heart Failure Group of Chinese Society of Cardiology, Editorial Board of Chinese Journal of Cardiology. Chinese expert consensus on the diagnosis and treatment of transthyretin cardiac amyoidosis[J]. Chinese Journal of Cardiology, 2021, 49(4): 324-332. | |
| [6] | Fontana M, Solomon S D, Kachadourian J, et al. CRISPR-Cas9 gene editing with nexiguran ziclumeran for ATTR cardiomyopathy[J]. N Engl J Med, 2024, 391(23): 2231-2241. |
| [7] | Gillmore J D, Solomon S D, Taubel J, et al. Updated phase 1 clinical trial outcomes of CRISPR gene editing with nexiguran ziclumeran in patients with transthyretin amyloidosis with cardiomyopathy[C]//Late-Breaking Science Abstracts and Featured Science Abstracts From the American Heart Association's Scientific Sessions 2025. Dallas: American Heart Association, 2025: e580-e581. |
| [8] | Dewey F E, Gusarova V, Dunbar R L, et al. Genetic and pharmacologic inactivation of ANGPTL3 and cardiovascular disease[J]. N Engl J Med, 2017, 377(3): 211-221. |
| [9] | Laffin L J, Nicholls S J, Scott R S, et al. Phase 1 trial of CRISPR-Cas9 gene editing targeting ANGPTL3[J]. N Engl J Med, 2025, 393(21): 2119-2130. |
| [10] | Bikard D, Jiang W Y, Samai P, et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system[J]. Nucleic Acids Res, 2013, 41(15): 7429-7437. |
| [11] | Gilbert L A, Horlbeck M A, Adamson B, et al. Genome-scale CRISPR-mediated control of gene repression and activation[J]. Cell, 2014, 159(3): 647-661. |
| [12] | Limpitikul W B, Dick I E, Tester D J, et al. A precision medicine approach to the rescue of function on malignant calmodulinopathic long-QT syndrome[J]. Circ Res, 2017, 120(1): 39-48. |
| [13] | Maeder M L, Linder S J, Cascio V M, et al. CRISPR RNA-guided activation of endogenous human genes[J]. Nat Methods, 2013, 10(10): 977-979. |
| [14] | Schoger E, Carroll K J, Iyer L M, et al. CRISPR-mediated activation of endogenous gene expression in the postnatal heart[J]. Circ Res, 2020, 126(1): 6-24. |
| [15] | Lauerer A M, Caravia X M, Maier L S, et al. Gene editing in common cardiovascular diseases[J]. Pharmacol Ther, 2024, 263: 108720. |
| [16] | Komor A C, Kim Y B, Packer M S, et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage[J]. Nature, 2016, 533(7603): 420-424. |
| [17] | Gaudelli N M, Komor A C, Rees H A, et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage[J]. Nature, 2017, 551(7681): 464-471. |
| [18] | Reichart D, Newby G A, Wakimoto H, et al. Efficient in vivo genome editing prevents hypertrophic cardiomyopathy in mice[J]. Nat Med, 2023, 29(2): 412-421. |
| [19] | Erickson J R, Joiner M L, Guan X Q, et al. A dynamic pathway for calcium-independent activation of CaMKⅡ by methionine oxidation[J]. Cell, 2008, 133(3): 462-474. |
| [20] | Lebek S, Chemello F, Caravia X M, et al. Ablation of CaMKⅡδ oxidation by CRISPR-Cas9 base editing as a therapy for cardiac disease[J]. Science, 2023, 379(6628): 179-185. |
| [21] | Lebek S, Caravia X M, Chemello F, et al. Elimination of CaMKⅡδ autophosphorylation by CRISPR-Cas9 base editing improves survival and cardiac function in heart failure in mice[J]. Circulation, 2023, 148(19): 1490-1504. |
| [22] | Bao X H, Liang Y J, Chang H M, et al. Targeting proprotein convertase subtilisin/kexin type 9 (PCSK9): from bench to bedside[J]. Signal Transduct Target Ther, 2024, 9(1): 13. |
| [23] | Lee R G, Mazzola A M, Braun M C, et al. Efficacy and safety of an investigational single-course CRISPR base-editing therapy targeting PCSK9 in nonhuman primate and mouse models[J]. Circulation, 2023, 147(3): 242-253. |
| [24] | Musunuru K, Chadwick A C, Mizoguchi T, et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates[J]. Nature, 2021, 593(7859): 429-434. |
| [25] | Vafai S B, Gladding P A, Scott R, et al. Safety and pharmacodynamic effects of VERVE-101, an investigational DNA base editing medicine designed to durably inactivate the PCSK9 gene and lower ldl cholesterol: interim results of the phase 1b heart-1 trial[C]//Late-Breaking Science Abstracts and Featured Science Abstracts From the American Heart Association′s Scientific Sessions 2023 and Late-Breaking Abstracts in Resuscitation Science From the Resuscitation Science Symposium 2023. Dallas: American Heart Association, 2023: e290-e291. |
| [26] | Nelson J W, Randolph P B, Shen S P, et al. Engineered pegRNAs improve prime editing efficiency[J]. Nat Biotechnol, 2022, 40(3): 402-410. |
| [27] | Anzalone A V, Randolph P B, Davis J R, et al. Search-and-replace genome editing without double-strand breaks or donor DNA[J]. Nature, 2019, 576(7785): 149-157. |
| [28] | Shuto Y, Nakagawa R, Zhu S Y, et al. Structural basis for pegRNA-guided reverse transcription by a prime editor[J]. Nature, 2024, 631(8019): 224-231. |
| [29] | Anzalone A V, Koblan L W, Liu D R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors[J]. Nat Biotechnol, 2020, 38(7): 824-844. |
| [30] | Gao P, Lyu Q, Ghanam A R, et al. Prime editing in mice reveals the essentiality of a single base in driving tissue-specific gene expression[J]. Genome Biol, 2021, 22(1): 83. |
| [31] | O′Leary K. Prime time for gene editing[J]. Nat Med, 2024, 30(12): 3392-3393. |
| [32] | Pickar-Oliver A, Gersbach C A. The next generation of CRISPR-Cas technologies and applications[J]. Nat Rev Mol Cell Biol, 2019, 20(8): 490-507. |
| [33] | Yang P, Lou Y M, Geng Z L, et al. Allele-specific suppression of variant MHC with high-precision RNA nuclease CRISPR-Cas13d prevents hypertrophic cardiomyopathy[J]. Circulation, 2024, 150(4): 283-298. |
| [34] | Cox D B T, Gootenberg J S, Abudayyeh O O, et al. RNA editing with CRISPR-cas13[J]. Science, 2017, 358(6366): 1019-1027. |
| [35] | Abudayyeh O O, Gootenberg J S, Franklin B, et al. A cytosine deaminase for programmable single-base RNA editing[J]. Science, 2019, 365(6451): 382-386. |
| [36] | Abudayyeh O O, Gootenberg J S, Essletzbichler P, et al. RNA targeting with CRISPR-cas13[J]. Nature, 2017, 550(7675): 280-284. |
| [37] | Booth B J, Nourreddine S, Katrekar D, et al. RNA editing: expanding the potential of RNA therapeutics[J]. Mol Ther, 2023, 31(6): 1533-1549. |
| [38] | YolTech Therapeutics Co, Ltd. A Phase Ⅰ/Ⅱa,open-label, single ascending dose and dose-expansion clinical study to evaluate the safety, tolerability, pharmacokinetics and pharmacodynamics of YOLT-201 in patients with transthyretin amyloidosis polyneuropathy (ATTR-PN) or transthyretin amyloidosis cardiomyopathy (ATTR-CM)[EB/OL]. (2024-08-06)[2026-01-25]. https://clinicaltrials.gov/study/NCT06539208. |
| [39] | Suzhou Dushu Lake Hospital. ART001 in the treatment of patients with transthyretin amyloidosis[EB/OL]. (2024-02-26) [2026-01-25]. https://www.chictr.org.cn/showprojEN.html?proj=210566. |
| [40] | Wan P, Tang S, Lin D, et al. Base editing gene therapy for heterozygous familial hypercholesterolemia[EB/OL]. (2025-04-17) [2026-01-26]. https://www.medrxiv.org/content/10.1101/2025.04.17. 25325983v1. |
| [41] | Verve Therapeutics Inc. Open-label, phase 1b, single ascending dose study to evaluate the safety of VERVE-102 administered to patients with heterozygous familial hypercholesterolemia or premature coronary artery disease who require additional lowering of low-density lipoprotein cholesterol[EB/OL]. (2025-12-17) [2026-01-25]. https://clinicaltrials.gov/study/NCT06164730. |
| [42] | CRISPR Therapeutics AG. A phase 1 open-label, multicenter, first-in-human, ascending single dose study evaluating the safety and tolerability of a lipid nanoparticle formulation of CRISPR-guide RNA-Cas9 nuclease (CTX320) for in vivo editing of the apolipoprotein(a) gene (LPA) in subjects with elevated lipoprotein(a) and a history of atherosclerotic cardiovascular disease or calcific aortic valve stenosis[EB/OL]. (2023-10-18) [2026-01-25]. https://anzctr.org.au/Trial/Registration/TrialReview.aspx?ACTRN=12623001095651. |
| [43] | Verve Therapeutics, Inc. A phase 1b single ascending dose study to evaluate the safety of VERVE-201 in patients with refractory hyperlipidemia[EB/OL]. (2025-11-28)[2026-01-25]. https://clinicaltrials.gov/study/NCT06451770. |
| [44] | Roberts A L, Connolly K L, Kirse D J, et al. Detection of group A Streptococcus in tonsils from pediatric patients reveals high rate of asymptomatic streptococcal carriage[J]. BMC Pediatr, 2012, 12: 3. |
| [45] | Lowy F D. Staphylococcus aureus infections[J]. N Engl J Med, 1998, 339(8): 520-532. |
| [46] | Charlesworth C T, Deshpande P S, Dever D P, et al. Identification of preexisting adaptive immunity to Cas9 proteins in humans[J]. Nat Med, 2019, 25(2): 249-254. |
| [47] | Raghavan R, Friedrich M J, King I, et al. Rational engineering of minimally immunogenic nucleases for gene therapy[J]. Nat Commun, 2025, 16(1): 105. |
| [48] | Doudna J A. The promise and challenge of therapeutic genome editing[J]. Nature, 2020, 578(7794): 229-236. |
| [49] | Ewaisha R, Anderson K S. Immunogenicity of CRISPR therapeutics: critical considerations for clinical translation[J]. Front Bioeng Biotechnol, 2023, 11: 1138596. |
| [50] | Jones S K Jr, Hawkins J A, Johnson N V, et al. Massively parallel kinetic profiling of natural and engineered CRISPR nucleases[J]. Nat Biotechnol, 2021, 39(1): 84-93. |
| [51] | Kim Y H, Kim N, Okafor I, et al. Sniper2L is a high-fidelity Cas9 variant with high activity[J]. Nat Chem Biol, 2023, 19(8): 972-980. |
| [52] | Zuo E W, Sun Y D, Wei W, et al. Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos[J]. Science, 2019, 364(6437): 289-292. |
| [53] | Grünewald J, Zhou R H, Garcia S P, et al. Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors[J]. Nature, 2019, 569(7756): 433-437. |
| [54] | Gehrke J M, Cervantes O, Clement M K, et al. An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities[J]. Nat Biotechnol, 2018, 36(10): 977-982. |
| [55] | Wessels H H, Méndez-Mancilla A, Guo X Y, et al. Massively parallel Cas13 screens reveal principles for guide RNA design[J]. Nat Biotechnol, 2020, 38(6): 722-727. |
| [56] | Liang S Q, Liu P P, Ponnienselvan K, et al. Genome-wide profiling of prime editor off-target sites in vitro and in vivo using PE-tag[J]. Nat Methods, 2023, 20(6): 898-907. |
| [57] | Tsai S Q, Zheng Z L, Nguyen N T, et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases[J]. Nat Biotechnol, 2015, 33(2): 187-197. |
| [58] | Tsai S Q, Nguyen N T, Malagon-Lopez J, et al. CIRCLE-seq: a highly sensitive in vitro screen for genome-wide CRISPR-Cas9 nuclease off-targets[J]. Nat Methods, 2017, 14(6): 607-614. |
| [59] | Bae, Park J, Kim J S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases[J]. Bioinformatics, 2014, 30(10): 1473-1475. |
| [60] | Ruffolo J A, Nayfach S, Gallagher J, et al. Design of highly functional genome editors by modelling CRISPR-Cas sequences[J]. Nature, 2025, 645(8080): 518-525. |
| [61] | Vo Q D. Gene editing therapy as a therapeutic approach for cardiovascular diseases in animal models: a scoping review[J]. PLoS One, 2025, 20(6): e0325330. |
| [62] | Wu S, Yang P, Geng Z L, et al. Base editing effectively prevents early-onset severe cardiomyopathy in Mybpc3 mutant mice[J]. Cell Res, 2024, 34(4): 327-330. |
| [63] | Böhm S, Splith V, Riedmayr L M, et al. A gene therapy for inherited blindness using dCas9-VPR-mediated transcriptional activation[J]. Sci Adv, 2020, 6(34): eaba5614. |
| [64] | Liu J J, Orlova N, Oakes B L, et al. CasX enzymes comprise a distinct family of RNA-guided genome editors[J]. Nature, 2019, 566(7743): 218-223. |
| [65] | Al-Shayeb B, Skopintsev P, Soczek K M, et al. Diverse virus-encoded CRISPR-Cas systems include streamlined genome editors[J]. Cell, 2022, 185(24): 4574-4586.e16. |
| [66] | Davis J R, Wang X, Witte I P, et al. Efficient in vivo base editing via single adeno-associated viruses with size-optimized genomes encoding compact adenine base editors[J]. Nat Biomed Eng, 2022, 6(11): 1272-1283. |
| [67] | Yang L Z, Liu Z Z, Chen G L, et al. microRNA-122-mediated liver detargeting enhances the tissue specificity of cardiac genome editing[J]. Circulation, 2024, 149(22): 1778-1781. |
| [68] | Kotterman M A, Schaffer D V. Engineering adeno-associated viruses for clinical gene therapy[J]. Nat Rev Genet, 2014, 15(7): 445-451. |
| [69] | Lisowski L, Dane A P, Chu K, et al. Selection and evaluation of clinically relevant AAV variants in a xenograft liver model[J]. Nature, 2014, 506(7488): 382-386. |
| [70] | Cullis P R, Felgner P L. The 60-year evolution of lipid nanoparticles for nucleic acid delivery[J]. Nat Rev Drug Discov, 2024, 23(9): 709-722. |
| [71] | Sun D, Lu Z R. Structure and function of cationic and ionizable lipids for nucleic acid delivery[J]. Pharm Res, 2023, 40(1): 27-46. |
| [72] | Chander N, Basha G, Yan Cheng M H, et al. Lipid nanoparticle mRNA systems containing high levels of sphingomyelin engender higher protein expression in hepatic and extra-hepatic tissues[J]. Mol Ther Methods Clin Dev, 2023, 30: 235-245. |
| [73] | Radmand A, Kim H, Beyersdorf J, et al. Cationic cholesterol-dependent LNP delivery to lung stem cells, the liver, and heart[J]. Proc Natl Acad Sci U S A, 2024, 121(11): e2307801120. |
| [74] | Yang J P, Yang Z L, Wang H, et al. A polymeric nanoparticle to co-deliver mitochondria-targeting peptides and Pt(Ⅳ) prodrug: toward high loading efficiency and combination efficacy[J]. Angew Chem Int Ed, 2024, 63(18): e202402291. |
| [75] | Yin D, Zhong Y Y, Ling S K, et al. Dendritic-cell-targeting virus-like particles as potent mRNA vaccine carriers[J]. Nat Biomed Eng, 2025, 9(2): 185-200. |
| [76] | Raguram A, An M R, Chen P Z, et al. Directed evolution of engineered virus-like particles with improved production and transduction efficiencies[J]. Nat Biotechnol, 2025, 43(10): 1635-1647. |
| [1] | GAO Xinjie, LIU Yan, WANG Dawei. Research progress and considerations for thalassemia gene therapy [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2025, 45(5): 540-548. |
| [2] | MA Huihua, YAN Kuipo, LIU Gang, XU Yazhou, ZHANG Lei, SUN Yanqin. Causal relationship between gut microbiota and cardiovascular diseases: a bidirectional Mendelian randomization analysis [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2025, 45(12): 1606-1619. |
| [3] | QIN Zihao, YUAN Hong, LU Yao, LENG Yiming. Multiple roles of CD34⁺ cell heterogeneity in cardiovascular injury and repair [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2025, 45(12): 1662-1670. |
| [4] | LUO Yuqi, WANG Yangyang, REN Jun. Advances in research on left ventricular hypertrophy-associated cardiovascular pathology in Alzheimer' s disease [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2025, 45(12): 1679-1686. |
| [5] | JIANG Lingyong. Status and advances in the mechanism research on dento-maxillofacial skeletal abnormalities [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2024, 44(6): 663-675. |
| [6] | LI Ping, JIANG Huiru, YE Mengyue, WANG Yayu, CHEN Xiaoyu, YUAN Ancai, XU Wenjie, DAI Huimin, CHEN Xi, YAN Xiaoxiang, TU Shengxian, ZHENG Yuanqi, ZHANG Wei, PU Jun. Analysis of epidemiological characteristics of risk factors for cardiovascular diseases and malignant tumors based on the Shanghai community elderly cohort [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2024, 44(5): 617-625. |
| [7] | GAO Xin, YANG Yiling, HUANG Xiangru, DAI Qinggang, JIANG Lingyong. Effect of Piezo1 on osteogenic differentiation of mouse bone marrow mesenchymal stem cells C3H10T1/2 based on CRISPR/Cas9 [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2023, 43(9): 1080-1088. |
| [8] | WANG Jingyi, WANG Qiong. Establishment of a mouse embryonic stem cell line carrying a reporter of mT-F2A-EGFP based on CRISPR/Cas9n technology [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2023, 43(4): 417-427. |
| [9] | ZHOU Yue, CHENG Chen, ZHENG Enlin, MENG Zhuo, WANG Jian, WANG Qingjie, HE Yongning, SUN Kun. Exploring potential new receptors for ELABELA in human embryonic stem cells by Crispr/Cas9-mediated gene editing system [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2022, 42(9): 1258-1264. |
| [10] | Xieyire·HAMULATI , ZHAO Qian, LI Cheng, SONG Ning, WANG Ying, Gulijiehere·TUERXUN , PU Jun, YANG Yining, LI Xiaomei. Clinical characteristics and health economics evaluation of real-world-based ischemic cardiovascular and cerebrovascular co-morbidities [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2022, 42(6): 778-785. |
| [11] | HAN Xiaxia, GU Shuangshuang, DAI Dai, SHEN Nan. Application of CRISPR/Cas9-mediated gene editing system to studying the regulation of T-bet in B cells [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2022, 42(4): 433-442. |
| [12] | HE Zhijie, HE Jinchun, ZHANG Yanpei, WANG Yaodong, WANG Zhanke. Analysis of lipoprotein subclasses of family with familial hypertriglyceridemia based on VAP method [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2022, 42(4): 482-489. |
| [13] | ZHANG Tong, TIAN Xue, ZUO Yingting, ZHENG Manqi, ZHANG Yijun, WU Shouling, CHEN Shuohua, MA Gaoting, TONG Xu, WANG Anxin, MO Dapeng. Association of triglyceride-glucose index with cardiovascular disease in people without traditional risk factors [J]. Journal of Shanghai Jiao Tong University (Medical Science), 2022, 42(3): 267-274. |
| [14] | Ming-qi CAI, Yan CHEN, Kai-bin LIN, Dong HUANG. Advances in the role of growth differentiation factor 11 in cardiovascular diseases [J]. JOURNAL OF SHANGHAI JIAOTONG UNIVERSITY (MEDICAL SCIENCE), 2021, 41(6): 834-838. |
| [15] | Yan-na ZHAO, Rong QIU, Nan SHEN, Yuan-jia TANG. Construction of inducible CRISPR/Cas9 system for studying gene function in mouse [J]. JOURNAL OF SHANGHAI JIAOTONG UNIVERSITY (MEDICAL SCIENCE), 2021, 41(3): 297-301. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||