NO是由不同的一氧化氮合酶(nitric oxide synthase,NOS)亚型产生的,这些亚型包括神经型NOS(NOS1)、诱导型NOS(NOS2)和内皮型NOS(NOS3)。每种亚型的表达具有独特的组织特异性,从而在不同的生理和病理条件下介导特定的NO的生物学功能。其中NOS1是神经系统中主要的NOS亚型,介导NO在神经系统中的生理效应[7-8]。既往研究[9-10]表明,NOS1的单核苷酸多态性(single nucleotide polymorphism,SNP)rs11615756增加人类日间小睡行为的发生概率(OR=1.018,95%CI 1.016~1.021,P=1.40×10-49),而其另一SNP rs7305526则与OSA患者最低血氧饱和度(lowest pulse blood oxygen saturation,LSpO2)呈显著正相关(OR=1.110,95%CI 1.071~1.149,P=1.41×10-8),提示NOS1基因可能在人类睡眠以及OSA临床数量性状调节中发挥关键作用。本研究拟利用英国生物银行(United Kingdom Biobank,UKB;
1 材料与方法
1.1 纳入数据
1.1.1 UKB GWAS数据集
本研究基于UKB不同时期GWAS数据集。通过检索欧洲生物信息学研究所(European Bioinformatics Institute,EMBL-EBI)GWAS数据库(
1.1.2 SSHS队列临床监测数据
SSHS队列的参与者来自2007年起就诊于上海交通大学医学院附属第六人民医院耳鼻咽喉头颈外科的OSA疑似患者。所有参与者都填写了全面的问卷调查,涉及主观睡眠质量、人口学信息、健康状况、家族病史、医疗史和生活方式因素(吸烟和饮酒)。主观睡眠质量由Epworth嗜睡量表(Epworth Sleepiness Scale,ESS)[16]和鼾声评分(Snoring Scale Score,SSS)[17]测评得出。睡眠数据采集自标准的多导睡眠监测(polysomnography,PSG)。PSG采集数据包括呼吸暂停低通气指数(apnea hypopnea index,AHI)、LSpO2、氧减指数(oxygen desaturation index,ODI)、血氧饱和度低于90%/80%/70%总时长(length of time with SpO2<90%/80%/70%,LT90/80/70)、总在床时间(time in bed,TIB)、睡眠周期时间(sleep period time,SPT)、总睡眠时间(total sleep time,TST)、非快速眼动期(non-rapid eye movement,NREM)、NREM 1/2/3期(N1/2/3)、快速眼动期(rapid eye movement,REM)、觉醒期(wakefulness,WK)。后续进一步计算TIB、SPT或TST期间REM、WK,以及N1、N2和N2期持续时间占比。所有参与者全基因组水平的SNP基因分型通过Affymetrix Genome-Wide Human SNP Array 6.0和Affymetrix AxiomTM Genome-Wide CHB Array芯片确定。
1.2 数据分析方法
1.2.1 UKB GWAS数据集分析
将获取的数据集标准化和统一化后进行后续分析。利用R软件(4.0.5版)及forestplot包根据计算出的OR、95%CI和P值绘制森林图,比较特定位点与性状的相关性。
1.2.2 SSHS临床监测数据分析
本研究共纳入SSHS中5 255例病例和764例对照的标准PSG数据。利用PLINK(1.90版)软件对基因型数据质量控制后,进行基因型与各OSA临床数量性状之间的线性回归分析,计算OR及其95%CI和P值。分析的OSA临床数量性状包括呼吸、血氧以及睡眠结构性状。利用LocusZoom(
1.3 人脑基因表达和表达数量性状位点分析
利用Allen Human Brain Atlas(AHBA)人脑基因表达数据库(
2 结果
2.1 NOS1 在人类全脑169个解剖结构的表达模式
图1
图1
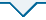
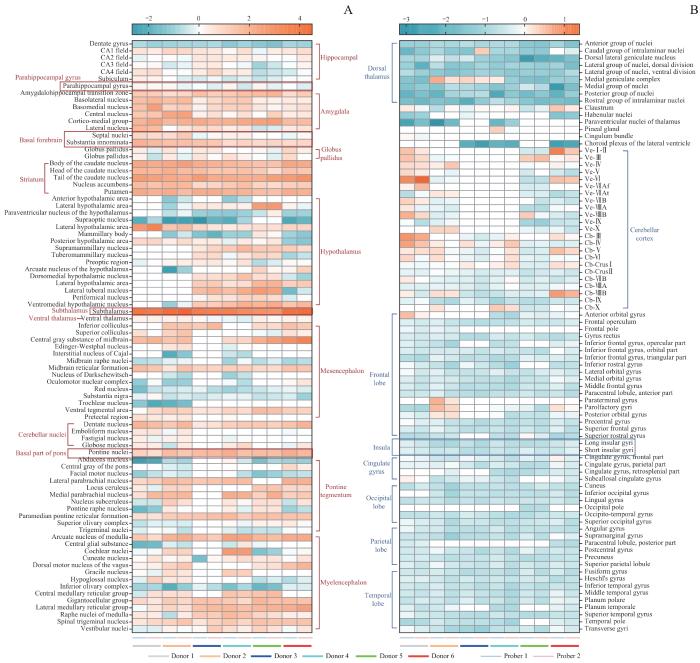
AHBA数据库6位捐赠者169个核团中 NOS1 基因表达情况热图
Note: A. Brain regions with relatively high levels of NOS1 gene expression. B. Brain regions with relatively low or no NOS1 expression.
Fig 1
Heatmap of the expression levels of the NOS1 gene in 169 nuclei from 6 donors in the AHBA database
2.2 基因功能注释
表1 rs7305526和rs11615756的eQTL分析
Tab 1
| SNP | Tissue | Effect size | P value |
|---|---|---|---|
| rs7305526 | Brain cortex | -0.127 | 0.044 |
| Lung | -0.136 | <0.001 | |
| Tibial nerve | 0.178 | <0.001 | |
| Colon sigmoid | -0.061 | 0.039 | |
| Heart atrial appendage | -0.155 | 0.014 | |
| Minor salivary gland | 0.141 | 0.033 | |
| Thyroid | 0.099 | 0.049 | |
| rs11615756 | Brain amygdala | -0.224 | 0.008 |
| Brain cortex | -0.151 | 0.028 | |
| Esophagus mucosa | -0.084 | 0.031 | |
| Thyroid | 0.104 | 0.049 | |
| Skeletal muscle | 0.061 | 0.017 | |
| Pancreas | 0.114 | 0.020 |
2.3 rs7305526和rs11615756与人类睡眠性状的关联
表2 本研究使用的UKB睡眠性状相关GWAS数据集
Tab 2
| Sleep trait | GWAS catalog accession | PMID | Author and year | Population | Number |
|---|---|---|---|---|---|
| Insomnia | GCST004695 | 28604731 | HAMMERSCHLAG 2017 | European (UK) | 113 006 |
| GCST90267286 | 37106081 | SCHOELER 2023 | European (UK) | 283 595 | |
| Sleep duration | GCST003839 | 27494321 | JONES 2016 | European (UK) | 127 573 |
| GCST006914 | 30531941 | DOHERTY 2018 | European (UK) | 91 105 | |
| GCST007561 | 30846698 | DASHTI 2019 | European (UK) | 446 118 |
图2
图2
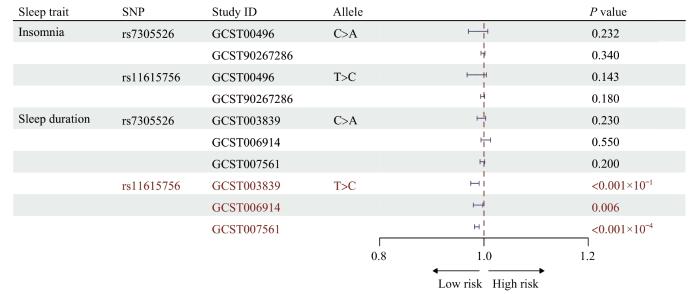
rs7305526和rs11615756与失眠和睡眠时长的相关性
Fig 2
Associations of genetic variants rs7305526 and rs11615756 with insomnia and sleep duration
2.4 rs7305526和rs11615756与OSA临床数量性状的关联
为深入探究NOS1与OSA发生发展和临床数量性状的关联,利用SSHS临床监测数据进行相关性研究。
2.4.1 OSA发病风险关联分析
图3
图3
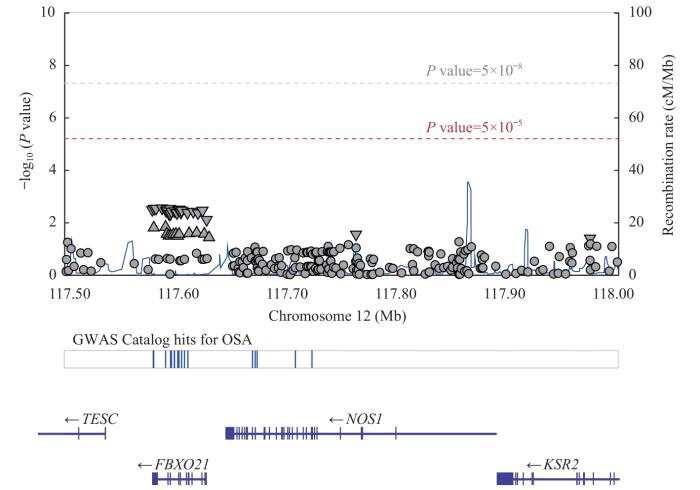
LocusZoom分析 NOS1 SNP与OSA发病风险的关联
Note: TESC—tescalcin; FBXO21—F-box protein 21; KSR2—kinase suppressor of RAS 2; Mb—megabase; cM—centimorgan.
Fig 3
LocusZoom analysis of the associations between NOS1 SNPs with OSA occurrence
表3 rs7305526各基因型特征
Tab 3
| Variable | Overall (n=6 026) | AA (n=1 941) | CA (n=2 867) | CC (n=1 077) | P value |
|---|---|---|---|---|---|
| Genotype/n(%) | <0.001 | ||||
| AA | 1 941 (32.2) | 1 941 (100.0) | 0 (0) | 0 (0) | |
| CA | 2 867 (47.6) | 0 (0) | 2 867 (100.0) | 0 (0) | |
| CC | 1 077 (17.9) | 0 (0) | 0 (0) | 1 077 (100.0) | |
| NA | 141 (2.3) | 0 (0) | 0 (0) | 0 (0) | |
| Gender/n(%) | 0.175 | ||||
| Male | 5 325 (88.4) | 1 699 (87.5) | 2 538 (88.5) | 967 (89.8) | |
| Female | 701 (11.6) | 242 (12.5) | 329 (11.5) | 110 (10.2) | |
| Smoking status/n(%) | 0.924 | ||||
| Never | 3 926 (65.2) | 1 262 (65.0) | 1 872 (65.3) | 708 (65.7) | |
| Ever | 2 100 (34.8) | 679 (35.0) | 995 (34.7) | 369 (34.3) | |
| Drinking status/n(%) | 0.862 | ||||
| Never | 3 637 (60.4) | 1 181 (60.8) | 1 722 (60.1) | 650 (60.4) | |
| Ever | 2 389 (39.6) | 760 (39.2) | 1 145 (39.9) | 427 (39.6) | |
| AHI (events/h)①/n(%) | <0.001 | ||||
| AHI< 5 (Non-OSA) | 764 (12.7) | 212 (10.9) | 360 (12.6) | 173 (16.1) | |
| 5 ≤AHI < 15 (Mild-OSA) | 318 (5.3) | 90 (4.6) | 161 (5.6) | 63 (5.9) | |
| 15 ≤AHI < 30 (Moderate-OSA) | 1 245 (20.7) | 407 (21.0) | 577 (20.1) | 235 (21.9) | |
| ≥30 (Severe-OSA) | 3 692 (61.3) | 1 231 (63.4) | 1 766 (61.6) | 603 (56.1) | |
| T2DM status/n(%) | 0.787 | ||||
| No | 5 580 (92.6) | 1 790 (92.2) | 2 658 (92.7) | 999 (92.8) | |
| Yes | 446 (7.4) | 151 (7.8) | 209 (7.3) | 78 (7.2) | |
| Hypertension status/n(%) | 0.761 | ||||
| No | 5 141 (85.3) | 1 649 (85.0) | 2 452 (85.5) | 925 (85.9) | |
| Yes | 885 (14.7) | 292 (15.0) | 415 (14.5) | 152 (14.1) |
表4 rs11615756各基因型特征
Tab 4
| Variable | Overall (n=6 026) | CC (n=3 352) | TC (n=2 236) | TT (n=385) | P value |
|---|---|---|---|---|---|
| Genotype/n(%) | <0.001 | ||||
| CC | 3 352 (55.6) | 3 352 (100.0) | 0 (0) | 0 (0) | |
| TC | 2 236 (37.1) | 0 (0) | 2 236 (100.0) | 0 (0) | |
| TT | 3 85 (6.4) | 0 (0) | 0 (0) | 385 (100.0) | |
| NA | 53 (0.9) | 0 (0) | 0 (0) | 0 (0) | |
| Gender/n(%) | 0.027 | ||||
| Male | 5 325 (88.4) | 2 965 (88.5) | 1 956 (87.5) | 355 (92.2) | |
| Female | 701 (11.6) | 387 (11.5) | 280 (12.5) | 30 (7.8) | |
| Smoking status/n(%) | 0.599 | ||||
| Never | 3 926 (65.2) | 2 183 (65.1) | 1 465 (65.5) | 242 (62.9) | |
| Ever | 2 100 (34.8) | 1 169 (34.9) | 771 (34.5) | 143 (37.1) | |
| Drinking status/n(%) | 0.513 | ||||
| Never | 3 637 (60.4) | 2 004 (59.8) | 1 371 (61.3) | 231 (60.0) | |
| Ever | 2 389 (39.6) | 1 348 (40.2) | 865 (38.7) | 154 (40.0) | |
| AHI (events/h)①/n(%) | 0.685 | ||||
| AHI< 5 (Non-OSA) | 764 (12.7) | 421 (12.6) | 293 (13.1) | 44 (11.4) | |
| 5 ≤AHI < 15 (Mild-OSA) | 318 (5.3) | 184 (5.5) | 109 (4.9) | 22 (5.7) | |
| 15 ≤AHI < 30 (Moderate-OSA) | 1 245 (20.7) | 700 (20.9) | 459 (20.6) | 70 (18.2) | |
| ≥30 (Severe-OSA) | 3 692 (61.3) | 2 044 (61.0) | 1 371 (61.4) | 249 (64.7) | |
| T2DM status/n(%) | 0.989 | ||||
| No | 5 580 (92.6) | 3 104 (92.6) | 2 072 (92.7) | 356 (92.5) | |
| Yes | 446 (7.4) | 248 (7.4) | 164 (7.3) | 29 (7.5) | |
| Hypertension status/n(%) | 0.568 | ||||
| No | 5 141 (85.3) | 2 863 (85.4) | 1 896 (84.8) | 334 (86.8) | |
| Yes | 885 (14.7) | 489 (14.6) | 340 (15.2) | 51 (13.2) |
表5 rs7305526和rs11615756在非OSA和OSA人群中的各基因型比率
Tab 5
| SNP | Genotype | Ratio of individuals of each genotype/% | |
|---|---|---|---|
| Non-OSA | OSA | ||
| rs7305526 | AA | 28.46 | 33.66 |
| CA | 48.32 | 48.78 | |
| CC | 23.22 | 17.55 | |
| rs11615756 | CC | 55.54 | 56.22 |
| TC | 38.65 | 37.23 | |
| TT | 5.80 | 6.55 | |
2.4.2 OSA呼吸、血氧和睡眠结构性状关联分析
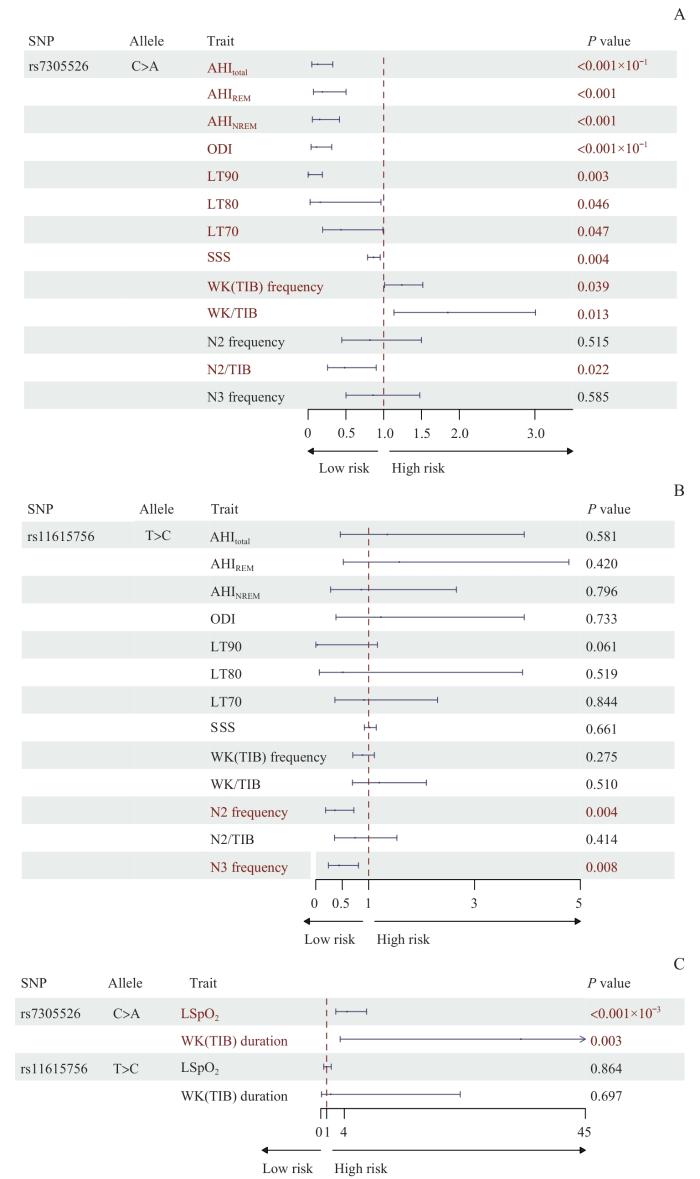
进一步使用加性模型分析rs7305526和rs11615756与OSA相关呼吸、血氧和睡眠结构性状的关系。结果:rs7305526(C>A)与部分OSA呼吸、血氧和睡眠性状显著相关。具体而言,在呼吸性状方面,rs7305526与总AHI(OR=0.126,95%CI 0.049~0.326,P<0.001)、AHIREM(OR=0.189,95%CI 0.071~0.502,P<0.001)、AHINREM(OR=0.153,95%CI 0.057~0.415,P<0.001)和SSS(OR=0.866,95%CI 0.787~0.954,P=0.004)呈显著负相关;在血氧性状方面,rs7305526同ODI(OR=0.111,95%CI 0.040~0.313,P<0.001)、LT90(OR=0.008,95%CI 0.001~0.188,P=0.003)、LT80(OR=0.160,95%CI 0.027~0.964,P=0.046)和LT70(OR=0.436,95%CI 0.192~0.990,P=0.047)呈显著负相关,与LSpO2(OR=4.455,95%CI 2.548~7.784,P<0.001)呈显著正相关;在睡眠结构性状方面,rs7305526与TIB中WK次数(OR=1.239,95%CI 1.011~1.517,P=0.039)、持续时长(OR=34.022,95%CI 3.300~350.724,P=0.003)和占比(OR=1.848,95%CI 1.136~3.007,P=0.013)呈显著正相关,而与N2占TIB的比例[(N2/TIB)%](OR=0.482,95%CI 0.258~0.901,P=0.022)呈显著负相关(图4A、C和表6)。而rs11615756(T>C)仅与睡眠结构性状中N2(OR=0.364,95%CI 0.184~0.720,P=0.004)和N3(OR=0.440,95%CI 0.240~0.807,P=0.008)次数呈显著负相关(图4B、C和表6),与其他呼吸、血氧及睡眠结构性状均无相关性。
图4
图4
rs7305526和rs11615756与OSA临床数量性状的关联
Note: A. Forest plot illustrating the association of genetic variant rs7305526 with clinical quantitative traits of OSA. B. Forest plot illustrating the associations of genetic variant rs11615756 with clinical quantitative traits of OSA. C. Forest plot illustrating the associations of rs7305526 and rs11615756 with additional clinical quantitative traits of OSA.
Fig 4
Associations of genetic variants rs7305526 and rs11615756 with clinical quantitative traits of OSA
表6 rs7305526和rs11615756与OSA临床数量性状的关联
Tab 6
| SNP | Allele | Trait | OR | 95%CI | P value | |
|---|---|---|---|---|---|---|
| Lower | Upper | |||||
| rs7305526 | C>A | AHItotal | 0.126 | 0.049 | 0.326 | <0.001 |
| AHIREM | 0.189 | 0.071 | 0.502 | <0.001 | ||
| AHINREM | 0.153 | 0.057 | 0.415 | <0.001 | ||
| ODI | 0.111 | 0.040 | 0.313 | <0.001 | ||
| LT90 | 0.008 | 0.001 | 0.188 | 0.003 | ||
| LT80 | 0.160 | 0.027 | 0.964 | 0.046 | ||
| LT70 | 0.436 | 0.192 | 0.990 | 0.047 | ||
| LSpO2 | 4.455 | 2.548 | 7.784 | <0.001 | ||
| SSS | 0.866 | 0.787 | 0.954 | 0.004 | ||
| WK(TIB) frequency | 1.239 | 1.011 | 1.517 | 0.039 | ||
| WK(TIB) duration | 34.022 | 3.300 | 350.724 | 0.003 | ||
| WK/TIB | 1.848 | 1.136 | 3.007 | 0.013 | ||
| N2 frequency | 0.818 | 0.446 | 1.500 | 0.515 | ||
| N2/TIB | 0.482 | 0.258 | 0.901 | 0.022 | ||
| N3 frequency | 0.861 | 0.501 | 1.477 | 0.586 | ||
| rs11615756 | T>C | AHItotal | 1.352 | 0.464 | 3.939 | 0.581 |
| AHIREM | 1.578 | 0.521 | 4.783 | 0.420 | ||
| AHINREM | 0.862 | 0.280 | 2.658 | 0.796 | ||
| ODI | 1.225 | 0.381 | 3.939 | 0.733 | ||
| LT90 | 0.032 | 0.001 | 1.165 | 0.061 | ||
| LT80 | 0.512 | 0.067 | 3.908 | 0.519 | ||
| LT70 | 0.911 | 0.361 | 2.301 | 0.844 | ||
| LSpO2 | 0.946 | 0.501 | 1.785 | 0.864 | ||
| SSS | 1.025 | 0.919 | 1.143 | 0.661 | ||
| WK(TIB) frequency | 0.880 | 0.700 | 1.107 | 0.275 | ||
| WK(TIB) duration | 1.690 | 0.120 | 23.712 | 0.697 | ||
| WK/TIB | 1.204 | 0.693 | 2.091 | 0.510 | ||
| N2 frequency | 0.364 | 0.184 | 0.720 | 0.004 | ||
| N2/TIB | 0.737 | 0.355 | 1.533 | 0.414 | ||
| N3 frequency | 0.440 | 0.240 | 0.807 | 0.008 | ||
2.4.3 OSA临床数量性状关联亚组分析
在女性中,NOS1 rs7305526和rs11615756与OSA临床数量性状均无明显相关性;而男性中,rs7305526与部分呼吸、血氧和睡眠结构性状的相关性同整体人群的趋势一致(表7)。根据AHI将入组患者划分为非OSA组,以及轻度、中度和重度OSA组。rs7305526与非OSA组(OR=2.039,95%CI 1.124~3.699,P=0.019)、中度OSA组(OR=2.280,95%CI 1.129~4.605,P=0.022)和重度OSA组(OR=2.217,95%CI 1.226~4.011,P=0.008)的LSpO2均呈显著正相关;然而在轻度OSA组,该位点仅与LT90(OR=23.117,95%CI 1.232~433.679,P=0.036)显著正相关。rs11615756与总人群的呼吸、血氧性状均无显著关联,而与非OSA组REM的AHI(OR=0.251,95%CI 0.074~0.856,P=0.027)和NREM的AHI(OR=0.270,95%CI 0.083~0.875,P=0.029)、轻度OSA组SSS(OR=0.626,95%CI 0.423~0.926,P=0.019)显著负相关,与中度OSA组LT70(OR=1.166,95%CI 1.011~1.344,P=0.035)显著正相关,与重度OSA组LT90(OR=0.002,95%CI 0.001~0.148,P=0.006)显著负相关(表8)。
表7 男性与女性rs7305526和rs11615756与OSA临床数量性状的关联
Tab 7
| Gender | SNP | Allele | Trait | OR | 95%CI | P value | |
|---|---|---|---|---|---|---|---|
| Lower | Upper | ||||||
| Male | rs7305526 | C>A | AHItotal | 0.103 | 0.037 | 0.285 | <0.001 |
| AHIREM | 0.189 | 0.066 | 0.535 | 0.002 | |||
| AHINREM | 0.111 | 0.038 | 0.322 | <0.001 | |||
| ODI | 0.093 | 0.031 | 0.281 | <0.001 | |||
| LT90 | 0.004 | 0.001 | 0.116 | 0.001 | |||
| LT80 | 0.177 | 0.029 | 1.087 | 0.061 | |||
| LT70 | 0.540 | 0.252 | 1.154 | 0.112 | |||
| LSpO2 | 4.782 | 2.821 | 8.108 | <0.001 | |||
| SSS | 0.865 | 0.781 | 0.958 | 0.006 | |||
| WK (TIB) frequency | 1.215 | 0.979 | 1.508 | 0.078 | |||
| WK (TIB) duration | 36.696 | 3.154 | 426.991 | 0.004 | |||
| WK/TIB | 2.002 | 1.208 | 3.318 | 0.007 | |||
| N2 frequency | 0.781 | 0.412 | 1.482 | 0.450 | |||
| N2/TIB | 0.129 | 0.006 | 2.867 | 0.195 | |||
| N3 frequency | 2.881 | 0.268 | 30.958 | 0.382 | |||
| rs11615756 | T>C | AHItotal | 1.251 | 0.398 | 3.929 | <0.001 | |
| AHIREM | 1.246 | 0.385 | 4.039 | 0.002 | |||
| AHINREM | 0.890 | 0.268 | 2.962 | <0.001 | |||
| ODI | 1.049 | 0.303 | 3.638 | <0.001 | |||
| LT90 | 0.049 | 0.001 | 2.051 | 0.001 | |||
| LT80 | 0.369 | 0.047 | 2.868 | 0.061 | |||
| LT70 | 0.748 | 0.317 | 1.764 | 0.112 | |||
| LSpO2 | 1.159 | 0.638 | 2.105 | <0.001 | |||
| SSS | 1.020 | 0.909 | 1.144 | 0.006 | |||
| WK (TIB) frequency | 0.862 | 0.676 | 1.100 | 0.078 | |||
| WK (TIB) duration | 4.212 | 0.264 | 67.122 | 0.004 | |||
| WK/TIB | 1.430 | 0.808 | 2.531 | 0.007 | |||
| N2 frequency | 0.356 | 0.174 | 0.726 | 0.450 | |||
| N2/TIB | 0.823 | 0.025 | 27.424 | 0.195 | |||
| N3 frequency | 0.197 | 0.014 | 2.876 | 0.382 | |||
| Female | rs7305526 | C>A | AHItotal | 0.726 | 0.060 | 8.786 | 0.801 |
| AHIREM | 0.371 | 0.025 | 5.499 | 0.471 | |||
| AHINREM | 1.586 | 0.106 | 23.735 | 0.738 | |||
| ODI | 0.761 | 0.045 | 12.926 | 0.850 | |||
| LT90 | 0.238 | 0.002 | 5 373.921 | 0.779 | |||
| LT80 | 0.128 | 0.001 | 195.286 | 0.581 | |||
| LT70 | 0.163 | 0.006 | 4.516 | 0.283 | |||
| LSpO2 | 1.137 | 0.266 | 4.854 | 0.862 | |||
| SSS | 0.846 | 0.635 | 1.127 | 0.253 | |||
| WK (TIB) frequency | 1.352 | 0.766 | 2.384 | 0.298 | |||
| WK (TIB) duration | 26.286 | 0.017 | 41 622.230 | 0.384 | |||
| WK/TIB | 1.857 | 0.394 | 8.760 | 0.434 | |||
| N2 frequency | 1.020 | 0.152 | 6.841 | 0.984 | |||
| N2/TIB | 0.002 | 0.001 | 2.374 | 0.078 | |||
| N3 frequency | 11.913 | 0.017 | 8 142.930 | 0.456 | |||
| rs11615756 | T>C | AHItotal | 3.274 | 0.174 | 61.738 | 0.428 | |
| AHIREM | 19.344 | 0.799 | 468.319 | 0.068 | |||
| AHINREM | 0.808 | 0.034 | 19.383 | 0.895 | |||
| ODI | 5.700 | 0.202 | 161.096 | 0.307 | |||
| LT90 | 0.004 | 0.001 | 716.665 | 0.375 | |||
| LT80 | 14.017 | 0.003 | 65 377.371 | 0.539 | |||
| LT70 | 20.663 | 0.444 | 960.811 | 0.122 | |||
| LSpO2 | 0.397 | 0.073 | 2.162 | 0.285 | |||
| SSS | 1.092 | 0.781 | 1.529 | 0.606 | |||
| WK (TIB) frequency | 1.047 | 0.537 | 2.040 | 0.893 | |||
| WK (TIB) duration | 0.002 | 0.001 | 3.107 | 0.088 | |||
| WK/TIB | 0.303 | 0.049 | 1.871 | 0.198 | |||
| N2 frequency | 0.546 | 0.057 | 5.229 | 0.599 | |||
| N2/TIB | 0.093 | 0.001 | 2 235.485 | 0.644 | |||
| N3 frequency | 0.096 | 0.001 | 201.306 | 0.547 | |||
表8 非OSA组,以及轻度、中度和重度OSA组rs7305526和rs11615756与OSA临床数量性状的关联
Tab 8
| SNP | Trait | Non-OSA (n=764) | Mild-OSA (n=318) | Moderate-OSA (n=1 245) | Severe-OSA (n=3 692) | ||||
|---|---|---|---|---|---|---|---|---|---|
P value | OR (95%CI) | P value | OR (95%CI) | P value | OR (95%CI) | P value | OR (95%CI) | ||
| rs7305526 | AHItotal | 0.530 | 0.955 (0.826‒1.104) | 0.290 | 0.818 (0.563‒1.188) | 0.619 | 1.090 (0.777‒1.528) | 0.206 | 0.593 (0.264‒1.334) |
| AHIREM | 0.763 | 1.176 (0.410‒3.380) | 0.965 | 1.070 (0.054‒21.232) | 0.890 | 0.894 (0.182‒4.383) | 0.577 | 0.749 (0.271‒2.070) | |
| AHINREM | 0.464 | 1.461 (0.529‒4.0332) | 0.143 | 4.276 (0.611‒29.910) | 0.846 | 1.076 (0.517‒2.239) | 0.206 | 0.538 (0.205‒1.408) | |
| ODI | 0.821 | 0.847 (0.199‒3.602) | 0.562 | 1.330 (0.505‒3.501) | 0.124 | 0.516 (0.222‒1.200) | 0.464 | 0.684 (0.247‒1.892) | |
| LT90 | 0.633 | 0.536 (0.041‒6.958) | 0.036 | 23.117 (1.232‒433.679) | 0.632 | 1.864 (0.146‒23.881) | 0.122 | 0.032 (0.001‒2.517) | |
| LT80 | 0.982 | 0.999 (0.907‒1.100) | 0.097 | 1.397 (0.941‒2.074) | 0.678 | 0.878 (0.475‒1.624) | 0.392 | 0.290 (0.017‒4.948) | |
| LT70 | 0.278 | 0.998 (0.993‒1.002) | 0.613 | 0.990 (0.951‒1.030) | 0.331 | 1.064 (0.939‒1.205) | 0.179 | 0.424 (0.121‒1.481) | |
| LSpO2 | 0.019 | 2.039 (1.124‒3.699) | 0.958 | 1.028 (0.372‒2.841) | 0.022 | 2.280 (1.129‒4.605) | 0.008 | 2.217 (1.226‒4.011) | |
| SSS | 0.421 | 0.888 (0.666‒1.186) | 0.333 | 1.193 (0.834‒1.706) | 0.995 | 1.001 (0.817‒1.226) | 0.153 | 0.923 (0.826‒1.030) | |
| WK (TIB) frequency | 0.215 | 1.498 (0.791‒2.838) | 0.016 | 0.369 (0.163‒0.832) | 0.130 | 1.432 (0.900‒2.281) | 0.244 | 1.158 (0.905‒1.4811) | |
| WK (TIB) duration | 0.119 | 406.376 (0.213‒775 370.391) | 0.606 | 0.060 (0.001‒2674.448) | 0.046 | 269.283 (1.095‒66 255.145) | 0.230 | 5.352 (0.345‒83.043) | |
| WK/TIB | 0.097 | 3.846 (0.784‒18.871) | 0.714 | 0.661 (0.072‒6.081) | 0.142 | 2.251 (0.763‒6.641) | 0.240 | 1.411 (0.794‒2.508) | |
| N2 frequency | 0.788 | 0.857 (0.277‒2.650 6) | 0.432 | 0.454 (0.063‒3.278) | 0.921 | 0.942 (0.289‒3.071) | 0.892 | 0.942 (0.398‒2.230) | |
| N2/TIB | 0.214 | 0.009 (0.001‒15.720) | 0.035 | 0.002 (0.001‒0.405) | 0.678 | 0.272 (0.001‒128.335) | 0.695 | 0.462 (0.010‒21.825) | |
| N3 frequency | 0.295 | 0.045 (0.001‒14.873) | 0.021 | 10 267.538 (3.963‒26 600 123.192) | 0.528 | 4.297 (0.046‒400.757) | 0.648 | 2.015 (0.099‒40.930) | |
| rs11615756 | AHItotal | 0.777 | 0.976 (0.823‒1.157) | 0.875 | 0.968 (0.639‒1.464) | 0.626 | 1.104 (0.742‒1.642) | 0.626 | 0.801 (0.328‒1.957) |
| AHIREM | 0.027 | 0.251 (0.074‒0.856) | 0.791 | 1.551 (0.060‒40.217) | 0.085 | 5.121 (0.800‒32.802) | 0.750 | 1.202 (0.389‒3.711) | |
| AHINREM | 0.029 | 0.270 (0.083‒0.875) | 0.504 | 0.500 (0.065‒3.838) | 0.392 | 1.457 (0.616‒3.448) | 0.377 | 0.620 (0.214‒1.793) | |
| ODI | 0.123 | 0.266 (0.050‒1.429) | 0.846 | 1.112 (0.381‒3.249) | 0.840 | 1.108 (0.411‒2.98) | 0.994 | 1.004 (0.327‒3.080) | |
| LT90 | 0.511 | 2.705 (0.139‒52.722) | 0.076 | 0.053 (0.002‒1.364) | 0.324 | 4.571 (0.222‒93.989) | 0.006 | 0.002 (0.001‒0.148) | |
| LT80 | 0.407 | 0.952 (0.848‒1.069) | 0.757 | 1.074 (0.682‒1.690) | 0.077 | 1.885 (0.933‒3.807) | 0.290 | 0.184 (0.008‒4.251) | |
| LT70 | 0.314 | 0.997 (0.992‒1.003) | 0.565 | 0.987 (0.943‒1.033) | 0.035 | 1.166 (1.011‒1.344) | 0.857 | 0.881 (0.221‒3.504) | |
| LSpO2 | 0.951 | 1.022 (0.511‒2.046) | 0.314 | 1.780 (0.577‒5.489) | 0.480 | 0.743 (0.326‒1.696) | 0.357 | 1.362 (0.706‒2.627) | |
| SSS | 0.697 | 1.070 (0.762‒1.501) | 0.019 | 0.626 (0.423‒0.926) | 0.949 | 0.992 (0.782‒1.259) | 0.435 | 1.050 (0.929‒1.186) | |
| WK (TIB) frequency | 0.238 | 0.636 (0.300‒1.349) | 0.835 | 1.101 (0.444‒2.731) | 0.917 | 0.971 (0.561‒1.682) | 0.470 | 0.905 (0.689‒1.188) | |
| WK (TIB) duration | 0.397 | 46.583 (0.006‒338 729.248) | 0.531 | 43.721 (0.001‒6 081 353.502) | 0.772 | 0.381 (0.001‒256.719) | 0.879 | 1.266 (0.061‒26.225) | |
| WK/TIB | 0.456 | 2.041 (0.313‒13.327) | 0.661 | 1.729 (0.148‒20.150) | 0.845 | 0.881 (0.245‒3.167) | 0.517 | 1.234 (0.653‒2.331) | |
| N2 frequency | 0.135 | 0.366 (0.098‒1.370) | 0.934 | 0.911 (0.099‒8.373) | 0.918 | 0.930 (0.234‒3.703) | 0.005 | 0.261 (0.101‒0.671) | |
| N2/TIB | 0.091 | 2 007.782 (0.296‒13 627 828.272) | 0.832 | 3.998 (0.001‒1 516 906.783) | 0.407 | 0.046 (0.001‒66.068) | 0.498 | 0.228 (0.003‒16.421) | |
| N3 frequency | 0.979 | 1.094 (0.001‒976.421) | 0.665 | 0.145 (0.001‒929.697) | 0.778 | 0.464 (0.002‒96.210) | 0.173 | 0.098 (0.004‒2.775) | |
3 讨 论
NO已被证实是一种促眠的气体信号分子。NOS1的表达水平和活性会直接影响中枢神经系统NO介导的生理功能,然而NOS1在人类睡眠调节以及OSA临床数量性状中的作用尚不清楚。我们研究发现,NOS1基因在人类睡眠和呼吸调控相关脑区、核团中高度表达,其遗传变异rs11615756与人类睡眠时长性状显著相关,另一个遗传变异rs7305526与OSA呼吸、血氧和睡眠结构等性状均呈显著相关,提示NOS1在人类睡眠时长以及OSA临床数量性状中起到重要的调节作用。
保护性位点NOS1 rs7305526(C>A)对AHI和SpO2产生了协同影响,这表明rs7305526遗传变异对血氧的调控并非独立存在,而是基于其引起的AHI的变化。尽管rs7305526(C>A)有助于改善OSA的呼吸和血氧性状,但会增加觉醒并缩短N2。这与我们所知道的OSA患者在通气改善后觉醒次数减少、NREM延长的情况相反。因此,rs7305526(C>A)对睡眠性状的调节独立于对呼吸性状的调节。
既往研究[21-23]表明,NO信号与脑衰老及相关的神经退行性疾病,如阿尔茨海默病、帕金森病和亨廷顿病密切相关。然而,具体的神经生物学机制仍待阐明。我们的研究发现NOS1遗传变异与睡眠时长性状显著相关,且NOS1定位于睡眠、呼吸相关脑区和认知记忆相关脑区——海马,而在大脑、小脑皮层中NOS1表达水平相对较低甚至不表达,提示NOS1可能更广泛地参与了睡眠和呼吸调节的过程,而并不深度参与躯体感觉、运动协调以及语言等神经生物学功能。海马区域负责清醒阶段记忆编码、睡眠阶段记忆重组,然而NOS1在海马以及睡眠调控区域均有高度表达,提示它可能通过调控睡眠和认知记忆形成过程进而影响阿尔茨海默病的发生[24-25]。
此前基于UKB的队列研究[9-10]发现NOS1 rs11615756遗传变异与日间小睡密切相关,而本研究同样基于UKB队列GWAS数据,发现该遗传变异与人类睡眠时长性状显著相关,然其在SSHS中仅与部分睡眠结构性状相关(N2和N3次数);此前基于SSHS发现的与OSA的LSpO2显著相关的位点rs7305526遗传变异与UKB队列的失眠和睡眠时长2种睡眠性状无显著相关性,但在SSHS中,该遗传变异不仅影响呼吸及血氧性状,还影响睡眠结构性状。NOS1不同遗传变异存在着不同的生物学效应,这种现象可能是由于NOS1的不同位点在调控性状时可能存在高度的人群特异性,也可能是由于不同位点受到不同信号的调节或表观遗传状态的影响。
NO供体药物已被广泛应用于不同的临床疾病治疗中。例如硝化甘油和硝普钠可通过稳定释放NO来松弛血管平滑肌,分别用于治疗高血压和心绞痛[26]。OSA患者常因慢性间歇性缺氧导致血氧分压降低,易引起全身多脏器的缺氧损伤,甚至导致高血压、冠心病等多系统的并发症。NOS1作为睡眠和呼吸中枢的双重调节因子,已被证实与OSA患者呼吸、血氧以及睡眠结构等性状存在不同程度的相关性,可作为治疗靶点,用于纠正患者睡眠中的通气不足和睡眠结构紊乱。
本研究尚存在以下不足之处。虽然本研究收集自SSHS的中国汉族人群中最大的OSA遗传样本库,然而前来就诊的多为重度OSA患者,且男性患者占比较大。同时本研究分析的NOS1的脑区表达模式源自未知种族的健康捐赠者的正常人脑,可能与中国汉族OSA患者脑中NOS1的表达情况存在差异。此外,在人类睡眠性状分析中,本研究仅选取了失眠和睡眠时长2个性状,仍需更多类别的睡眠障碍数据集以评估NOS1在特定睡眠阶段中的作用。
NOS1基因rs7305526和rs11615756遗传变异与人类睡眠性状及OSA呼吸、血氧事件和睡眠结构等临床数量性状密切相关,该基因有望成为纠正OSA患者通气障碍和睡眠结构紊乱的重要治疗靶点。
作者贡献声明
袁灏琳、李念念和刘峰参与数据分析,袁灏琳、李念念、胡珺晖、沈锦虹、高振飞、关建、刘峰和殷善开参与论文写作和修改。
AUTHOR's CONTRIBUTIONS
YUAN Haolin, LI Niannian and LIU Feng participated in data analysis. YUAN Haolin, LI Niannian, HU Junhui, SHEN Jinhong, GAO Zhenfei, GUAN Jian, LIU Feng and YIN Shankai participated in manuscript drafting and revision.
利益冲突声明
所有作者声明不存在利益冲突。
COMPETING INTERESTS
All authors disclose no relevant conflict of interests.
参考文献