
上海交通大学学报(医学版) ›› 2026, Vol. 46 ›› Issue (3): 358-367.doi: 10.3969/j.issn.1674-8115.2026.03.010
• 综述 • 上一篇
收稿日期:2025-07-27
接受日期:2025-12-30
出版日期:2026-03-28
发布日期:2026-03-30
通讯作者:
闫小响,教授,研究员,副主任医师,博士;电子信箱:cardexyanxx@hotmail.com。基金资助:
Tian Qijia, Jia Kangni, Yan Xiaoxiang( )
)
Received:2025-07-27
Accepted:2025-12-30
Online:2026-03-28
Published:2026-03-30
Contact:
Yan Xiaoxiang, E-mail: cardexyanxx@hotmail.com.Supported by:摘要:
基因编辑技术可对生物体基因组或转录产物进行定点修饰。10余年来,以CRISPR/Cas9为代表的新一代基因编辑技术迅速发展,在效率、安全性和应用广度方面均优于传统锌指核酸酶(zinc finger nuclease,ZFN)和转录激活因子样效应物核酸酶(transcription activator-like effector nuclease,TALEN)技术,逐步从实验室研究走向临床转化阶段。心血管疾病是全球死亡和健康损害的主要原因之一,现有治疗手段常面临靶点难成药、副作用大及疗效有限等挑战。对于部分传统药物治疗无效的心血管疾病病例,基因编辑技术提供了新的治疗思路。目前,该技术已应用于针对转甲状腺素蛋白淀粉样变、高脂血症等疾病的药物开发,其中NTLA-2001和CTX310等药物已率先进入临床试验阶段。降低免疫反应与脱靶风险、突破心脏靶向递送瓶颈,是该技术实现临床转化的关键。随着人工智能辅助设计与非病毒载体等新技术的成熟,CRISPR/Cas技术有望突破当前的肝脏靶向限制,实现心脏原位治疗,为心血管疾病提供更具根治潜力的治疗策略。该文系统梳理CRISPR/Cas系统的结构与作用机制,总结近年来其在心血管疾病中的应用进展,包括基因敲除、表观编辑、碱基编辑、先导编辑及RNA编辑等多个方向,以期为该领域的后续研究与临床转化提供参考。
中图分类号:
田启嘉, 贾康妮, 闫小响. CRISPR/Cas基因编辑技术在心血管疾病中的应用研究进展[J]. 上海交通大学学报(医学版), 2026, 46(3): 358-367.
Tian Qijia, Jia Kangni, Yan Xiaoxiang. Advances in CRISPR/Cas gene editing and its applications in cardiovascular diseases[J]. Journal of Shanghai Jiao Tong University (Medical Science), 2026, 46(3): 358-367.
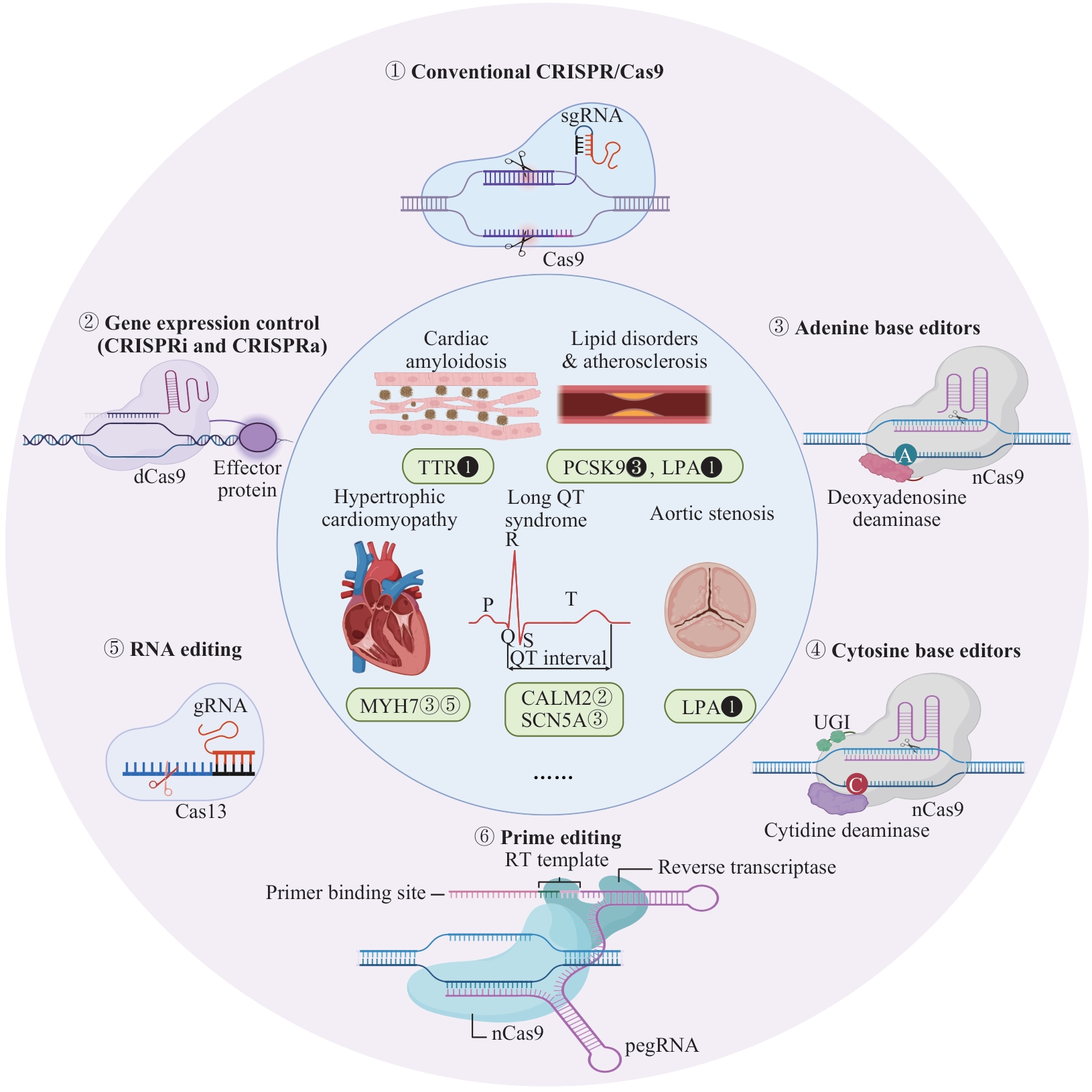
图1 CRISPR/Cas技术在心血管疾病中的应用示意图
Fig 1 Schematic diagram of CRISPR/Cas applications in cardiovascular diseasesNote: Outer ring: 6 main categories of gene editing tools; inner ring: associated cardiovascular diseases and therapeutic targets. Markers adjacent to gene targets indicate the corresponding editing technologies (matching the outer ring numbering). Solid markers denote therapies in clinical trials; hollow markers indicate therapies supported only by in vivo animal evidence. CALM2—calmodulin 2; CRISPRi/a—CRISPR interference/activation; dCas9—dead Cas9; LPA—lipoprotein(a); MYH7—myosin heavy chain 7; nCas9—Cas9 nickase; PCSK9—proprotein convertase subtilisin/kexin type 9; pegRNA—prime editing guide RNA; RT—reverse transcription; SCN5A—sodium voltage-gated channel alpha subunit 5; TTR—transthyretin; UGI—uracil glycosylase inhibitor.
| Treatment | Gene editing method | Indication | Target gene | Delivery method | Clinical trial ID | Phase | Efficacy | Follow-up | Safety | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| NTLA-2001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06128629 | Ⅲ | TTR reduction: 89% (Day 28), 90% (Month 12) | 87% reduction sustained (Month 36) | Most common treatment-related AEs: mild infusion-related reactions (14%) and transient liver enzyme elevations (6%) | [ |
| YOLT-201 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06539208 | Ⅰ/Ⅱa | Trial ongoing | \ | \ | [ |
| ART001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | ChiCTR2400081216 | Ⅰ | Trial ongoing | \ | \ | [ |
| YOLT-101 | Base editing | Familial hypercholesterolemia | PCSK9 | GalNAc-LNP | NCT06458010 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 72.5% and LDL-C by 50.4% (Month 4) | Follow-up ongoing | Most common treatment-related AEs; mild infusion-related reactions (83%) and transient liver enzyme elevations (50%) | [ |
| VERVE-101 | Base editing | Familial hypercholesterolemia | PCSK9 | LNP | NCT05398029 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 47% and LDL-C by 55% (Day 28); sustained through 180 d | Follow-up ongoing | Treatment-related Grade 3 serious AEs: transient liver enzyme elevations and thrombocytopenia; mild infusion-related reactions were common; trial enrollment paused due to safety concerns | [ |
| VERVE-102 | Base editing | Familial hypercholesterolemia or premature coronary artery disease | PCSK9 | GalNAc-LNP | NCT06164730 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX320 | CRISPR/Cas9 knock out | Atherosclerosis or calcific aortic valve stenosis | LPA | LNP | ACTRN12623001095651 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX310 | CRISPR/Cas9 knock out | Refractory dyslipidemias | ANGPTL3 | LNP | ACTRN12623000809639 | Ⅰ | High-dose (0.8 mg/kg): ANGPTL3 reduced by 73.2%, LDL-C by 48.9%, and triglycerides by 55.2% (Day 60) | Follow-up ongoing | Most common treatment-related AEs: mild infusion-related reactions (20%) and transient liver enzyme elevations (7%) | [ |
| VERVE-201 | Base editing | Refractory hypercholesterolemia | ANGPTL3 | GalNAc-LNP | NCT06451770 | Ⅰ | Trial ongoing | \ | \ | [ |
表1 已进入临床试验的心血管疾病领域基因编辑药物
Tab 1 Gene editing therapies for cardiovascular diseases in clinical trials
| Treatment | Gene editing method | Indication | Target gene | Delivery method | Clinical trial ID | Phase | Efficacy | Follow-up | Safety | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| NTLA-2001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06128629 | Ⅲ | TTR reduction: 89% (Day 28), 90% (Month 12) | 87% reduction sustained (Month 36) | Most common treatment-related AEs: mild infusion-related reactions (14%) and transient liver enzyme elevations (6%) | [ |
| YOLT-201 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | NCT06539208 | Ⅰ/Ⅱa | Trial ongoing | \ | \ | [ |
| ART001 | CRISPR/Cas9 knock out | ATTR-CM | TTR | LNP | ChiCTR2400081216 | Ⅰ | Trial ongoing | \ | \ | [ |
| YOLT-101 | Base editing | Familial hypercholesterolemia | PCSK9 | GalNAc-LNP | NCT06458010 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 72.5% and LDL-C by 50.4% (Month 4) | Follow-up ongoing | Most common treatment-related AEs; mild infusion-related reactions (83%) and transient liver enzyme elevations (50%) | [ |
| VERVE-101 | Base editing | Familial hypercholesterolemia | PCSK9 | LNP | NCT05398029 | Ⅰ | High-dose (0.6 mg/kg): PCSK9 reduced by 47% and LDL-C by 55% (Day 28); sustained through 180 d | Follow-up ongoing | Treatment-related Grade 3 serious AEs: transient liver enzyme elevations and thrombocytopenia; mild infusion-related reactions were common; trial enrollment paused due to safety concerns | [ |
| VERVE-102 | Base editing | Familial hypercholesterolemia or premature coronary artery disease | PCSK9 | GalNAc-LNP | NCT06164730 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX320 | CRISPR/Cas9 knock out | Atherosclerosis or calcific aortic valve stenosis | LPA | LNP | ACTRN12623001095651 | Ⅰ | Trial ongoing | \ | \ | [ |
| CTX310 | CRISPR/Cas9 knock out | Refractory dyslipidemias | ANGPTL3 | LNP | ACTRN12623000809639 | Ⅰ | High-dose (0.8 mg/kg): ANGPTL3 reduced by 73.2%, LDL-C by 48.9%, and triglycerides by 55.2% (Day 60) | Follow-up ongoing | Most common treatment-related AEs: mild infusion-related reactions (20%) and transient liver enzyme elevations (7%) | [ |
| VERVE-201 | Base editing | Refractory hypercholesterolemia | ANGPTL3 | GalNAc-LNP | NCT06451770 | Ⅰ | Trial ongoing | \ | \ | [ |
| [1] | Mensah G A, Fuster V, Murray C J L, et al. Global burden of cardiovascular diseases and risks, 1990‒2022[J]. J Am Coll Cardiol, 2023, 82(25): 2350-2473. |
| [2] | Kim Y G, Cha J, Chandrasegaran S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain[J]. Proc Natl Acad Sci U S A, 1996, 93(3): 1156-1160. |
| [3] | Christian M, Cermak T, Doyle E L, et al. Targeting DNA double-strand breaks with TAL effector nucleases[J]. Genetics, 2010, 186(2): 757-761. |
| [4] | Jinek M, Chylinski K, Fonfara I, et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity[J]. Science, 2012, 337(6096): 816-821. |
| [5] | 中华医学会心血管病学分会心力衰竭学组, 中华心血管病杂志编辑委员会. 转甲状腺素蛋白心脏淀粉样变诊断与治疗中国专家共识[J]. 中华心血管病杂志, 2021, 49(4): 324-332. |
| Heart Failure Group of Chinese Society of Cardiology, Editorial Board of Chinese Journal of Cardiology. Chinese expert consensus on the diagnosis and treatment of transthyretin cardiac amyoidosis[J]. Chinese Journal of Cardiology, 2021, 49(4): 324-332. | |
| [6] | Fontana M, Solomon S D, Kachadourian J, et al. CRISPR-Cas9 gene editing with nexiguran ziclumeran for ATTR cardiomyopathy[J]. N Engl J Med, 2024, 391(23): 2231-2241. |
| [7] | Gillmore J D, Solomon S D, Taubel J, et al. Updated phase 1 clinical trial outcomes of CRISPR gene editing with nexiguran ziclumeran in patients with transthyretin amyloidosis with cardiomyopathy[C]//Late-Breaking Science Abstracts and Featured Science Abstracts From the American Heart Association's Scientific Sessions 2025. Dallas: American Heart Association, 2025: e580-e581. |
| [8] | Dewey F E, Gusarova V, Dunbar R L, et al. Genetic and pharmacologic inactivation of ANGPTL3 and cardiovascular disease[J]. N Engl J Med, 2017, 377(3): 211-221. |
| [9] | Laffin L J, Nicholls S J, Scott R S, et al. Phase 1 trial of CRISPR-Cas9 gene editing targeting ANGPTL3[J]. N Engl J Med, 2025, 393(21): 2119-2130. |
| [10] | Bikard D, Jiang W Y, Samai P, et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system[J]. Nucleic Acids Res, 2013, 41(15): 7429-7437. |
| [11] | Gilbert L A, Horlbeck M A, Adamson B, et al. Genome-scale CRISPR-mediated control of gene repression and activation[J]. Cell, 2014, 159(3): 647-661. |
| [12] | Limpitikul W B, Dick I E, Tester D J, et al. A precision medicine approach to the rescue of function on malignant calmodulinopathic long-QT syndrome[J]. Circ Res, 2017, 120(1): 39-48. |
| [13] | Maeder M L, Linder S J, Cascio V M, et al. CRISPR RNA-guided activation of endogenous human genes[J]. Nat Methods, 2013, 10(10): 977-979. |
| [14] | Schoger E, Carroll K J, Iyer L M, et al. CRISPR-mediated activation of endogenous gene expression in the postnatal heart[J]. Circ Res, 2020, 126(1): 6-24. |
| [15] | Lauerer A M, Caravia X M, Maier L S, et al. Gene editing in common cardiovascular diseases[J]. Pharmacol Ther, 2024, 263: 108720. |
| [16] | Komor A C, Kim Y B, Packer M S, et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage[J]. Nature, 2016, 533(7603): 420-424. |
| [17] | Gaudelli N M, Komor A C, Rees H A, et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage[J]. Nature, 2017, 551(7681): 464-471. |
| [18] | Reichart D, Newby G A, Wakimoto H, et al. Efficient in vivo genome editing prevents hypertrophic cardiomyopathy in mice[J]. Nat Med, 2023, 29(2): 412-421. |
| [19] | Erickson J R, Joiner M L, Guan X Q, et al. A dynamic pathway for calcium-independent activation of CaMKⅡ by methionine oxidation[J]. Cell, 2008, 133(3): 462-474. |
| [20] | Lebek S, Chemello F, Caravia X M, et al. Ablation of CaMKⅡδ oxidation by CRISPR-Cas9 base editing as a therapy for cardiac disease[J]. Science, 2023, 379(6628): 179-185. |
| [21] | Lebek S, Caravia X M, Chemello F, et al. Elimination of CaMKⅡδ autophosphorylation by CRISPR-Cas9 base editing improves survival and cardiac function in heart failure in mice[J]. Circulation, 2023, 148(19): 1490-1504. |
| [22] | Bao X H, Liang Y J, Chang H M, et al. Targeting proprotein convertase subtilisin/kexin type 9 (PCSK9): from bench to bedside[J]. Signal Transduct Target Ther, 2024, 9(1): 13. |
| [23] | Lee R G, Mazzola A M, Braun M C, et al. Efficacy and safety of an investigational single-course CRISPR base-editing therapy targeting PCSK9 in nonhuman primate and mouse models[J]. Circulation, 2023, 147(3): 242-253. |
| [24] | Musunuru K, Chadwick A C, Mizoguchi T, et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates[J]. Nature, 2021, 593(7859): 429-434. |
| [25] | Vafai S B, Gladding P A, Scott R, et al. Safety and pharmacodynamic effects of VERVE-101, an investigational DNA base editing medicine designed to durably inactivate the PCSK9 gene and lower ldl cholesterol: interim results of the phase 1b heart-1 trial[C]//Late-Breaking Science Abstracts and Featured Science Abstracts From the American Heart Association′s Scientific Sessions 2023 and Late-Breaking Abstracts in Resuscitation Science From the Resuscitation Science Symposium 2023. Dallas: American Heart Association, 2023: e290-e291. |
| [26] | Nelson J W, Randolph P B, Shen S P, et al. Engineered pegRNAs improve prime editing efficiency[J]. Nat Biotechnol, 2022, 40(3): 402-410. |
| [27] | Anzalone A V, Randolph P B, Davis J R, et al. Search-and-replace genome editing without double-strand breaks or donor DNA[J]. Nature, 2019, 576(7785): 149-157. |
| [28] | Shuto Y, Nakagawa R, Zhu S Y, et al. Structural basis for pegRNA-guided reverse transcription by a prime editor[J]. Nature, 2024, 631(8019): 224-231. |
| [29] | Anzalone A V, Koblan L W, Liu D R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors[J]. Nat Biotechnol, 2020, 38(7): 824-844. |
| [30] | Gao P, Lyu Q, Ghanam A R, et al. Prime editing in mice reveals the essentiality of a single base in driving tissue-specific gene expression[J]. Genome Biol, 2021, 22(1): 83. |
| [31] | O′Leary K. Prime time for gene editing[J]. Nat Med, 2024, 30(12): 3392-3393. |
| [32] | Pickar-Oliver A, Gersbach C A. The next generation of CRISPR-Cas technologies and applications[J]. Nat Rev Mol Cell Biol, 2019, 20(8): 490-507. |
| [33] | Yang P, Lou Y M, Geng Z L, et al. Allele-specific suppression of variant MHC with high-precision RNA nuclease CRISPR-Cas13d prevents hypertrophic cardiomyopathy[J]. Circulation, 2024, 150(4): 283-298. |
| [34] | Cox D B T, Gootenberg J S, Abudayyeh O O, et al. RNA editing with CRISPR-cas13[J]. Science, 2017, 358(6366): 1019-1027. |
| [35] | Abudayyeh O O, Gootenberg J S, Franklin B, et al. A cytosine deaminase for programmable single-base RNA editing[J]. Science, 2019, 365(6451): 382-386. |
| [36] | Abudayyeh O O, Gootenberg J S, Essletzbichler P, et al. RNA targeting with CRISPR-cas13[J]. Nature, 2017, 550(7675): 280-284. |
| [37] | Booth B J, Nourreddine S, Katrekar D, et al. RNA editing: expanding the potential of RNA therapeutics[J]. Mol Ther, 2023, 31(6): 1533-1549. |
| [38] | YolTech Therapeutics Co, Ltd. A Phase Ⅰ/Ⅱa,open-label, single ascending dose and dose-expansion clinical study to evaluate the safety, tolerability, pharmacokinetics and pharmacodynamics of YOLT-201 in patients with transthyretin amyloidosis polyneuropathy (ATTR-PN) or transthyretin amyloidosis cardiomyopathy (ATTR-CM)[EB/OL]. (2024-08-06)[2026-01-25]. https://clinicaltrials.gov/study/NCT06539208. |
| [39] | Suzhou Dushu Lake Hospital. ART001 in the treatment of patients with transthyretin amyloidosis[EB/OL]. (2024-02-26) [2026-01-25]. https://www.chictr.org.cn/showprojEN.html?proj=210566. |
| [40] | Wan P, Tang S, Lin D, et al. Base editing gene therapy for heterozygous familial hypercholesterolemia[EB/OL]. (2025-04-17) [2026-01-26]. https://www.medrxiv.org/content/10.1101/2025.04.17. 25325983v1. |
| [41] | Verve Therapeutics Inc. Open-label, phase 1b, single ascending dose study to evaluate the safety of VERVE-102 administered to patients with heterozygous familial hypercholesterolemia or premature coronary artery disease who require additional lowering of low-density lipoprotein cholesterol[EB/OL]. (2025-12-17) [2026-01-25]. https://clinicaltrials.gov/study/NCT06164730. |
| [42] | CRISPR Therapeutics AG. A phase 1 open-label, multicenter, first-in-human, ascending single dose study evaluating the safety and tolerability of a lipid nanoparticle formulation of CRISPR-guide RNA-Cas9 nuclease (CTX320) for in vivo editing of the apolipoprotein(a) gene (LPA) in subjects with elevated lipoprotein(a) and a history of atherosclerotic cardiovascular disease or calcific aortic valve stenosis[EB/OL]. (2023-10-18) [2026-01-25]. https://anzctr.org.au/Trial/Registration/TrialReview.aspx?ACTRN=12623001095651. |
| [43] | Verve Therapeutics, Inc. A phase 1b single ascending dose study to evaluate the safety of VERVE-201 in patients with refractory hyperlipidemia[EB/OL]. (2025-11-28)[2026-01-25]. https://clinicaltrials.gov/study/NCT06451770. |
| [44] | Roberts A L, Connolly K L, Kirse D J, et al. Detection of group A Streptococcus in tonsils from pediatric patients reveals high rate of asymptomatic streptococcal carriage[J]. BMC Pediatr, 2012, 12: 3. |
| [45] | Lowy F D. Staphylococcus aureus infections[J]. N Engl J Med, 1998, 339(8): 520-532. |
| [46] | Charlesworth C T, Deshpande P S, Dever D P, et al. Identification of preexisting adaptive immunity to Cas9 proteins in humans[J]. Nat Med, 2019, 25(2): 249-254. |
| [47] | Raghavan R, Friedrich M J, King I, et al. Rational engineering of minimally immunogenic nucleases for gene therapy[J]. Nat Commun, 2025, 16(1): 105. |
| [48] | Doudna J A. The promise and challenge of therapeutic genome editing[J]. Nature, 2020, 578(7794): 229-236. |
| [49] | Ewaisha R, Anderson K S. Immunogenicity of CRISPR therapeutics: critical considerations for clinical translation[J]. Front Bioeng Biotechnol, 2023, 11: 1138596. |
| [50] | Jones S K Jr, Hawkins J A, Johnson N V, et al. Massively parallel kinetic profiling of natural and engineered CRISPR nucleases[J]. Nat Biotechnol, 2021, 39(1): 84-93. |
| [51] | Kim Y H, Kim N, Okafor I, et al. Sniper2L is a high-fidelity Cas9 variant with high activity[J]. Nat Chem Biol, 2023, 19(8): 972-980. |
| [52] | Zuo E W, Sun Y D, Wei W, et al. Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos[J]. Science, 2019, 364(6437): 289-292. |
| [53] | Grünewald J, Zhou R H, Garcia S P, et al. Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors[J]. Nature, 2019, 569(7756): 433-437. |
| [54] | Gehrke J M, Cervantes O, Clement M K, et al. An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities[J]. Nat Biotechnol, 2018, 36(10): 977-982. |
| [55] | Wessels H H, Méndez-Mancilla A, Guo X Y, et al. Massively parallel Cas13 screens reveal principles for guide RNA design[J]. Nat Biotechnol, 2020, 38(6): 722-727. |
| [56] | Liang S Q, Liu P P, Ponnienselvan K, et al. Genome-wide profiling of prime editor off-target sites in vitro and in vivo using PE-tag[J]. Nat Methods, 2023, 20(6): 898-907. |
| [57] | Tsai S Q, Zheng Z L, Nguyen N T, et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases[J]. Nat Biotechnol, 2015, 33(2): 187-197. |
| [58] | Tsai S Q, Nguyen N T, Malagon-Lopez J, et al. CIRCLE-seq: a highly sensitive in vitro screen for genome-wide CRISPR-Cas9 nuclease off-targets[J]. Nat Methods, 2017, 14(6): 607-614. |
| [59] | Bae, Park J, Kim J S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases[J]. Bioinformatics, 2014, 30(10): 1473-1475. |
| [60] | Ruffolo J A, Nayfach S, Gallagher J, et al. Design of highly functional genome editors by modelling CRISPR-Cas sequences[J]. Nature, 2025, 645(8080): 518-525. |
| [61] | Vo Q D. Gene editing therapy as a therapeutic approach for cardiovascular diseases in animal models: a scoping review[J]. PLoS One, 2025, 20(6): e0325330. |
| [62] | Wu S, Yang P, Geng Z L, et al. Base editing effectively prevents early-onset severe cardiomyopathy in Mybpc3 mutant mice[J]. Cell Res, 2024, 34(4): 327-330. |
| [63] | Böhm S, Splith V, Riedmayr L M, et al. A gene therapy for inherited blindness using dCas9-VPR-mediated transcriptional activation[J]. Sci Adv, 2020, 6(34): eaba5614. |
| [64] | Liu J J, Orlova N, Oakes B L, et al. CasX enzymes comprise a distinct family of RNA-guided genome editors[J]. Nature, 2019, 566(7743): 218-223. |
| [65] | Al-Shayeb B, Skopintsev P, Soczek K M, et al. Diverse virus-encoded CRISPR-Cas systems include streamlined genome editors[J]. Cell, 2022, 185(24): 4574-4586.e16. |
| [66] | Davis J R, Wang X, Witte I P, et al. Efficient in vivo base editing via single adeno-associated viruses with size-optimized genomes encoding compact adenine base editors[J]. Nat Biomed Eng, 2022, 6(11): 1272-1283. |
| [67] | Yang L Z, Liu Z Z, Chen G L, et al. microRNA-122-mediated liver detargeting enhances the tissue specificity of cardiac genome editing[J]. Circulation, 2024, 149(22): 1778-1781. |
| [68] | Kotterman M A, Schaffer D V. Engineering adeno-associated viruses for clinical gene therapy[J]. Nat Rev Genet, 2014, 15(7): 445-451. |
| [69] | Lisowski L, Dane A P, Chu K, et al. Selection and evaluation of clinically relevant AAV variants in a xenograft liver model[J]. Nature, 2014, 506(7488): 382-386. |
| [70] | Cullis P R, Felgner P L. The 60-year evolution of lipid nanoparticles for nucleic acid delivery[J]. Nat Rev Drug Discov, 2024, 23(9): 709-722. |
| [71] | Sun D, Lu Z R. Structure and function of cationic and ionizable lipids for nucleic acid delivery[J]. Pharm Res, 2023, 40(1): 27-46. |
| [72] | Chander N, Basha G, Yan Cheng M H, et al. Lipid nanoparticle mRNA systems containing high levels of sphingomyelin engender higher protein expression in hepatic and extra-hepatic tissues[J]. Mol Ther Methods Clin Dev, 2023, 30: 235-245. |
| [73] | Radmand A, Kim H, Beyersdorf J, et al. Cationic cholesterol-dependent LNP delivery to lung stem cells, the liver, and heart[J]. Proc Natl Acad Sci U S A, 2024, 121(11): e2307801120. |
| [74] | Yang J P, Yang Z L, Wang H, et al. A polymeric nanoparticle to co-deliver mitochondria-targeting peptides and Pt(Ⅳ) prodrug: toward high loading efficiency and combination efficacy[J]. Angew Chem Int Ed, 2024, 63(18): e202402291. |
| [75] | Yin D, Zhong Y Y, Ling S K, et al. Dendritic-cell-targeting virus-like particles as potent mRNA vaccine carriers[J]. Nat Biomed Eng, 2025, 9(2): 185-200. |
| [76] | Raguram A, An M R, Chen P Z, et al. Directed evolution of engineered virus-like particles with improved production and transduction efficiencies[J]. Nat Biotechnol, 2025, 43(10): 1635-1647. |
| [1] | 高欣洁, 刘艳, 王大威. 地中海贫血基因治疗研究进展及思考[J]. 上海交通大学学报(医学版), 2025, 45(5): 540-548. |
| [2] | 马会华, 闫奎坡, 刘刚, 徐亚洲, 张磊, 孙彦琴. 肠道微生物群与心血管疾病的因果关系评价:双向孟德尔随机化分析[J]. 上海交通大学学报(医学版), 2025, 45(12): 1606-1619. |
| [3] | 秦梓豪, 袁洪, 陆瑶, 冷一铭. CD34+细胞异质性在心血管损伤与修复中的多重角色[J]. 上海交通大学学报(医学版), 2025, 45(12): 1662-1670. |
| [4] | 罗宇琦, 王洋洋, 任骏. 左心室肥厚相关心血管病变影响阿尔茨海默病的研究进展[J]. 上海交通大学学报(医学版), 2025, 45(12): 1679-1686. |
| [5] | 江凌勇. 牙颌面骨畸形机制研究的现状与发展[J]. 上海交通大学学报(医学版), 2024, 44(6): 663-675. |
| [6] | 李萍, 蒋惠如, 叶梦月, 王雅玉, 陈潇雨, 袁安彩, 徐文杰, 戴慧敏, 陈曦, 闫小响, 涂圣贤, 郑元琦, 张薇, 卜军. 基于上海社区老年人群队列的心血管疾病和恶性肿瘤的危险因素流行特征分析[J]. 上海交通大学学报(医学版), 2024, 44(5): 617-625. |
| [7] | 高昕, 杨屹羚, 黄湘如, 代庆刚, 江凌勇. 利用CRISPR/Cas9靶向敲除Piezo1基因对小鼠间充质干细胞成骨分化的影响研究[J]. 上海交通大学学报(医学版), 2023, 43(9): 1080-1088. |
| [8] | 王靖怡, 王琼. 基于CRISPR/Cas9n技术建立携带mT-F2A-EGFP报告系统的小鼠胚胎干细胞系[J]. 上海交通大学学报(医学版), 2023, 43(4): 417-427. |
| [9] | 周月, 程晨, 郑恩霖, 孟卓, 王鉴, 王青洁, 何勇宁, 孙锟. 利用Crispr/Cas9基因编辑系统在人胚胎干细胞中探索ELABELA的潜在新受体[J]. 上海交通大学学报(医学版), 2022, 42(9): 1258-1264. |
| [10] | 韩夏夏, 顾霜霜, 戴黛, 沈南. 应用CRISPR/Cas9介导的基因编辑系统研究B细胞中转录因子T-bet的调控作用[J]. 上海交通大学学报(医学版), 2022, 42(4): 433-442. |
| [11] | 何志洁, 何津春, 张燕培, 王耀东, 王占科. 基于垂直密度梯度离心全自动血脂谱检测法的家族性高三酰甘油血症家系血脂亚组分分析[J]. 上海交通大学学报(医学版), 2022, 42(4): 482-489. |
| [12] | 张彤, 田雪, 左颖婷, 郑曼琪, 张怡君, 吴寿岭, 陈朔华, 马高亭, 佟旭, 王安心, 莫大鹏. 无传统危险因素人群中TyG指数与心脑血管疾病的关系[J]. 上海交通大学学报(医学版), 2022, 42(3): 267-274. |
| [13] | 蔡明琪, 陈焱, 林开斌, 黄冬. 生长分化因子11在心血管疾病中的作用[J]. 上海交通大学学报(医学版), 2021, 41(6): 834-838. |
| [14] | 赵艳娜, 邱荣, 沈南, 唐元家. 构建诱导型CRISPR/Cas9系统用于小鼠免疫细胞基因功能研究[J]. 上海交通大学学报(医学版), 2021, 41(3): 297-301. |
| [15] | 孙敏, 张冬颖. 钠-葡萄糖共转运蛋白2抑制剂对2型糖尿病患者心血管保护作用的研究进展[J]. 上海交通大学学报(医学版), 2021, 41(3): 391-395. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||