
上海交通大学学报(医学版) ›› 2026, Vol. 46 ›› Issue (4): 451-466.doi: 10.3969/j.issn.1674-8115.2026.04.005
• 论著 · 基础研究 • 上一篇
收稿日期:2025-09-25
接受日期:2025-12-23
出版日期:2026-04-15
发布日期:2026-04-15
通讯作者:
肖 辉,主任医师,博士;电子信箱:xiaohui771210@163.com。基金资助:
Peng Qianqian, Song Jinghan, Xu Xingyi, Xiao Hui( )
)
Received:2025-09-25
Accepted:2025-12-23
Online:2026-04-15
Published:2026-04-15
Contact:
Xiao Hui, E-mail: xiaohui771210@163.com.Supported by:摘要:
目的·系统鉴定RNA结合蛋白人类抗原R(human antigen R,HuR)在非小细胞肺癌(non-small cell lung cancer,NSCLC)中的下游关键靶基因,并阐明其通过调控该靶点影响肿瘤进展的分子机制。方法·基于癌症基因组图谱(The Cancer Genome Atlas,TCGA)数据库、基因型-组织表达项目(Genotype-Tissue Expression,GTEx)数据库及临床蛋白质组学肿瘤分析联盟(Clinical Proteomic Tumor Analysis Consortium,CPTAC)数据库,分析HuR在NSCLC及癌旁组织中的表达差异;通过免疫组织化学法检测HuR蛋白水平,并利用GSE19188数据集进行验证。基于TCGA数据,分析HuR表达与临床病理参数间关系,进行生存分析,并通过单因素与多因素Cox回归评估预后因素,构建列线图模型预测生存率。利用ESTIMATE包及CIBERSORT算法分析HuR表达与免疫微环境的相关性。采用Transwell实验检测HuR敲低后A549细胞迁移与侵袭能力变化。通过RNA测序筛选差异表达基因(differentially expressed genes,DEGs),结合蛋白质相互作用(protein-protein interaction,PPI)网络鉴定枢纽基因,并利用基因本体论(Gene Ontology,GO)与京都基因与基因组百科全书(Kyoto Encyclopedia of Genes and Genomes,KEGG)进行富集分析。采用RNA免疫沉淀(RNA immunoprecipitation,RIP)验证HuR与整合素β1(integrin β 1,ITGB1) mRNA的直接结合,并通过实时定量PCR(RT-qPCR)和Western blotting分别检测HuR敲低后ITGB1在mRNA和蛋白水平的表达变化。结果·HuR在肺癌组织中显著高表达,与不良预后独立相关(P=0.045),且其表达与免疫抑制性微环境特征呈负相关。功能实验表明敲低HuR可显著抑制肺癌细胞的迁移(P<0.001)与侵袭(P=0.002)能力。生物信息学分析将ITGB1锁定为HuR下游的核心枢纽基因,富集分析显示其参与细胞外基质受体相互作用等通路。RIP实验证实HuR可直接结合ITGB1 mRNA;进一步的RT-qPCR与Western blotting结果表明,敲低HuR导致ITGB1在mRNA和蛋白水平均显著下调,提示HuR主要通过维持ITGB1 mRNA稳定性发挥转录后调控作用。结论·HuR通过直接结合并稳定ITGB1 mRNA,激活下游信号通路,促进NSCLC进展。HuR-ITGB1调控轴的发现不仅为深入理解肺癌的发病机制提供了新视角,也为预后判断和靶向治疗提供了潜在新靶点。
中图分类号:
彭倩倩, 宋璟涵, 徐杏怡, 肖辉. RNA结合蛋白HuR通过调控ITGB1促进非小细胞肺癌进展的生物信息学分析及验证[J]. 上海交通大学学报(医学版), 2026, 46(4): 451-466.
Peng Qianqian, Song Jinghan, Xu Xingyi, Xiao Hui. Bioinformatic analysis and validation of the RNA-binding protein HuR promoting non-small cell lung cancer progression via ITGB1[J]. Journal of Shanghai Jiao Tong University (Medical Science), 2026, 46(4): 451-466.
| TCGA | Detail | Tumor/n | Normal/n | GTEx | Normal/n |
|---|---|---|---|---|---|
| ACC | Adrenocortical carcinoma | 79 | ‒ | Adrenal gland | 128 |
| BLCA | Bladder urothelial carcinoma | 408 | 19 | Bladder | 9 |
| BRCA | Breast invasive carcinoma | 1 093 | 112 | Breast | 179 |
| CESC | Cervical squamous cell carcinoma and endocervical adenocarcinoma | 304 | 3 | Cervix uteri | 10 |
| CHOL | Cholangio carcinoma | 36 | 9 | ‒ | ‒ |
| COAD | Colon adenocarcinoma | 457 | 41 | Colon | 308 |
| DLBC | Lymphoid neoplasm diffuse large B-cell lymphoma | 48 | ‒ | Blood | 337 |
| ESCA | Esophageal carcinoma | 184 | 11 | Esophagus | 273 |
| GBM | Glioblastoma multiforme | 153 | 5 | Brain | 207 |
| HNSC | Head and neck squamous cell carcinoma | 520 | 44 | ‒ | ‒ |
| KICH | Kidney chromophobe | 66 | 25 | Kidney | 28 |
| KIRC | Kidney renal clear cell carcinoma | 533 | 72 | Kidney | 28 |
| KIRP | Kidney renal papillary cell carcinoma | 290 | 32 | Kidney | 28 |
| LAML | Acute myeloid leukemia | 173 | ‒ | Bone marrow | 70 |
| LGG | Brain lower grade glioma | 516 | ‒ | Brain | 207 |
| LIHC | Liver hepatocellular carcinoma | 371 | 50 | Liver | 110 |
| LUAD | Lung adenocarcinoma | 515 | 59 | Lung | 288 |
| LUSC | Lung squamous cell carcinoma | 501 | 50 | Lung | 288 |
| MESO | Mesothelioma | 87 | ‒ | ‒ | ‒ |
| OV | Ovarian serous cystadenocarcinoma | 303 | ‒ | Ovary | 88 |
| PAAD | Pancreatic adenocarcinoma | 178 | 4 | Pancreas | 167 |
| PCPG | Pheochromocytoma and paraganglioma | 179 | 3 | ‒ | ‒ |
| PRAD | Prostate adenocarcinoma | 497 | 52 | Prostate | 100 |
| READ | Rectum adenocarcinoma | 166 | 10 | Colon | 308 |
| SARC | Sarcoma | 259 | 2 | ‒ | ‒ |
| SKCM | Skin cutaneous melanoma | 103 | ‒ | Skin | 557 |
| STAD | Stomach adenocarcinoma | 415 | 35 | Stomach | 175 |
| TGCT | Testicular germ cell tumor | 150 | ‒ | Testis | 165 |
| THCA | Thyroid carcinoma | 501 | 59 | Thyroid | 278 |
| THYM | Thymoma | 120 | ‒ | Blood | 337 |
| UCEC | Uterine corpus endometrial carcinoma | 545 | 35 | Uterus | 78 |
| UCS | Uterine carcinosarcoma | 57 | ‒ | Uterus | 78 |
| UVM | Uveal melanoma | 80 | ‒ | ‒ | ‒ |
| Total | 9 887 | 732 | 3 555 |
表1 TCGA与GTEx数据集肿瘤及正常组织样本数
Tab 1 Sample counts of tumor and normal tissues from TCGA and GTEx datasets
| TCGA | Detail | Tumor/n | Normal/n | GTEx | Normal/n |
|---|---|---|---|---|---|
| ACC | Adrenocortical carcinoma | 79 | ‒ | Adrenal gland | 128 |
| BLCA | Bladder urothelial carcinoma | 408 | 19 | Bladder | 9 |
| BRCA | Breast invasive carcinoma | 1 093 | 112 | Breast | 179 |
| CESC | Cervical squamous cell carcinoma and endocervical adenocarcinoma | 304 | 3 | Cervix uteri | 10 |
| CHOL | Cholangio carcinoma | 36 | 9 | ‒ | ‒ |
| COAD | Colon adenocarcinoma | 457 | 41 | Colon | 308 |
| DLBC | Lymphoid neoplasm diffuse large B-cell lymphoma | 48 | ‒ | Blood | 337 |
| ESCA | Esophageal carcinoma | 184 | 11 | Esophagus | 273 |
| GBM | Glioblastoma multiforme | 153 | 5 | Brain | 207 |
| HNSC | Head and neck squamous cell carcinoma | 520 | 44 | ‒ | ‒ |
| KICH | Kidney chromophobe | 66 | 25 | Kidney | 28 |
| KIRC | Kidney renal clear cell carcinoma | 533 | 72 | Kidney | 28 |
| KIRP | Kidney renal papillary cell carcinoma | 290 | 32 | Kidney | 28 |
| LAML | Acute myeloid leukemia | 173 | ‒ | Bone marrow | 70 |
| LGG | Brain lower grade glioma | 516 | ‒ | Brain | 207 |
| LIHC | Liver hepatocellular carcinoma | 371 | 50 | Liver | 110 |
| LUAD | Lung adenocarcinoma | 515 | 59 | Lung | 288 |
| LUSC | Lung squamous cell carcinoma | 501 | 50 | Lung | 288 |
| MESO | Mesothelioma | 87 | ‒ | ‒ | ‒ |
| OV | Ovarian serous cystadenocarcinoma | 303 | ‒ | Ovary | 88 |
| PAAD | Pancreatic adenocarcinoma | 178 | 4 | Pancreas | 167 |
| PCPG | Pheochromocytoma and paraganglioma | 179 | 3 | ‒ | ‒ |
| PRAD | Prostate adenocarcinoma | 497 | 52 | Prostate | 100 |
| READ | Rectum adenocarcinoma | 166 | 10 | Colon | 308 |
| SARC | Sarcoma | 259 | 2 | ‒ | ‒ |
| SKCM | Skin cutaneous melanoma | 103 | ‒ | Skin | 557 |
| STAD | Stomach adenocarcinoma | 415 | 35 | Stomach | 175 |
| TGCT | Testicular germ cell tumor | 150 | ‒ | Testis | 165 |
| THCA | Thyroid carcinoma | 501 | 59 | Thyroid | 278 |
| THYM | Thymoma | 120 | ‒ | Blood | 337 |
| UCEC | Uterine corpus endometrial carcinoma | 545 | 35 | Uterus | 78 |
| UCS | Uterine carcinosarcoma | 57 | ‒ | Uterus | 78 |
| UVM | Uveal melanoma | 80 | ‒ | ‒ | ‒ |
| Total | 9 887 | 732 | 3 555 |
| CPTAC | Detail | Tumor/n | Normal/n |
|---|---|---|---|
| BRCA | Breast cancer | 126 | 18 |
| CCRCC | Clear cell renal cell carcinoma | 117 | 84 |
| COAD | Colon adenocarcinoma | 101 | 96 |
| GBM | Glioblastoma | 156 | 5 |
| HNSC | Head and neck squamous cell carcinoma | 110 | 68 |
| NSCLC | Non-small cell lung cancer | 217 | 199 |
| OV | Ovarian serous cystadenocarcinoma | 93 | 10 |
| UCEC | Uterine corpus endometrial carcinoma | 104 | 30 |
| PAAD | Pancreatic adenocarcinoma | 137 | 74 |
| LIHC | Liver hepatocellular carcinoma | 128 | 106 |
| Total | 1 289 | 690 |
表2 CPTAC数据集肿瘤与配对正常组织样本数
Tab 2 Sample counts of tumor and matched normal tissues from the CPTAC dataset
| CPTAC | Detail | Tumor/n | Normal/n |
|---|---|---|---|
| BRCA | Breast cancer | 126 | 18 |
| CCRCC | Clear cell renal cell carcinoma | 117 | 84 |
| COAD | Colon adenocarcinoma | 101 | 96 |
| GBM | Glioblastoma | 156 | 5 |
| HNSC | Head and neck squamous cell carcinoma | 110 | 68 |
| NSCLC | Non-small cell lung cancer | 217 | 199 |
| OV | Ovarian serous cystadenocarcinoma | 93 | 10 |
| UCEC | Uterine corpus endometrial carcinoma | 104 | 30 |
| PAAD | Pancreatic adenocarcinoma | 137 | 74 |
| LIHC | Liver hepatocellular carcinoma | 128 | 106 |
| Total | 1 289 | 690 |
| Gene | Forward (5'→3') | Reverse (5'→3') |
|---|---|---|
| HuR | TAATCGCCATAGCCTTCCTAA | GGCGTCTGCAAATGGTTGTA |
| β-actin | ATCAAGATCATTGCTCCTCCTGAG | CTGCTTGCTGATCCACATCTG |
| ITGB1 | ATCCCAGAGGCTCCAAGAT | CCCCTGATCTTAATCGCAA |
表3 RT-qPCR引物序列
Tab 3 Primer sequences for RT-qPCR
| Gene | Forward (5'→3') | Reverse (5'→3') |
|---|---|---|
| HuR | TAATCGCCATAGCCTTCCTAA | GGCGTCTGCAAATGGTTGTA |
| β-actin | ATCAAGATCATTGCTCCTCCTGAG | CTGCTTGCTGATCCACATCTG |
| ITGB1 | ATCCCAGAGGCTCCAAGAT | CCCCTGATCTTAATCGCAA |
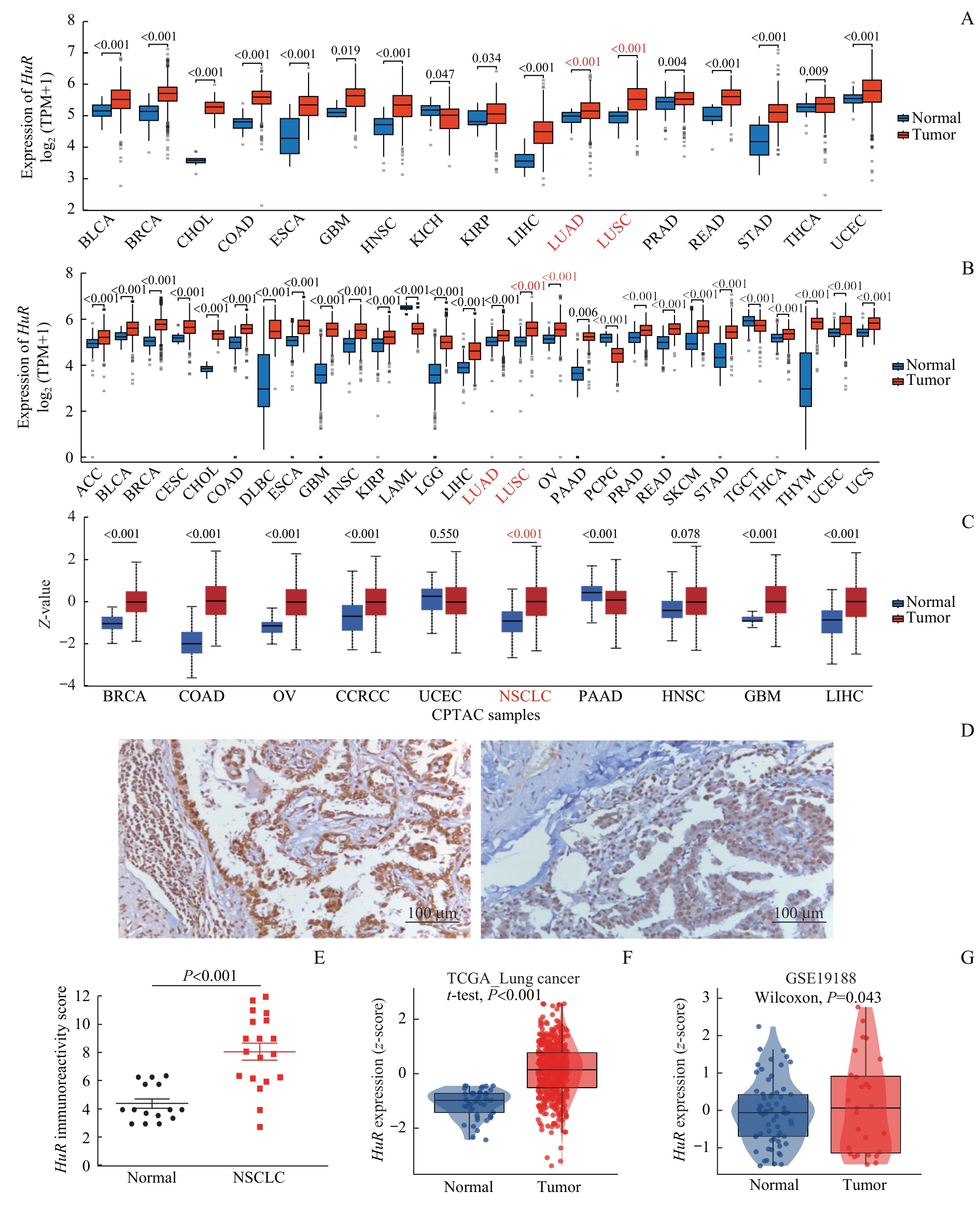
图1 肺癌组织中 HuR 的上调表达Note: A/B. Expression of HuR in 33 cancer types compared with normal tissues based on TCGA (A) and TCGA combined with the GTEx (B) datasets, with only statistically significant comparisons displayed. C. Differential expression of HuR between tumor and matched normal tissues across 10 cancer types from the CTPAC database. D. Representative immunohistochemistry images of HuR protein in lung cancer and adjacent normal tissues. E. Immunoreactivity scores of HuR in lung cancer versus normal tissues. F/G. Validation of HuR overexpression in lung cancer using TCGA (F) and GSE19188 (G) datasets. ACC—adrenocortical carcinoma; BLCA—bladder urothelial carcinoma; BRCA—breast invasive carcinoma; CESC—cervical squamous cell carcinoma and endocervical adenocarcinoma; CHOL—cholangiocarcinoma; COAD—colon adenocarcinoma; DLBC—lymphoid neoplasm diffuse large B-cell lymphoma; ESCA—esophageal carcinoma; GBM—glioblastoma multiforme; HNSC—head and neck squamous cell carcinoma; KIRP—kidney renal papillary cell carcinoma; LAML—acute myeloid leukemia; LGG—brain lower grade glioma; LIHC—liver hepatocellular carcinoma; LUAD—lung adenocarcinoma; LUSC—lung squamous cell carcinoma; OV—ovarian serous cystadenocarcinoma; PAAD—pancreatic adenocarcinoma; PCPG—pheochromocytoma and paraganglioma; PRAD—prostate adenocarcinoma; READ—rectum adenocarcinoma; STAD—stomach adenocarcinoma; TGCT—testicular germ cell tumor; THCA—thyroid carcinoma; THYM—thymoma; UCEC—Uterine corpus endometrial carcinoma; UCS—uterine carcinosarcoma; CCRCC—clear cell renal cell carcinoma.
Fig 1 Up-regulated expression of HuR in lung cancer tissues
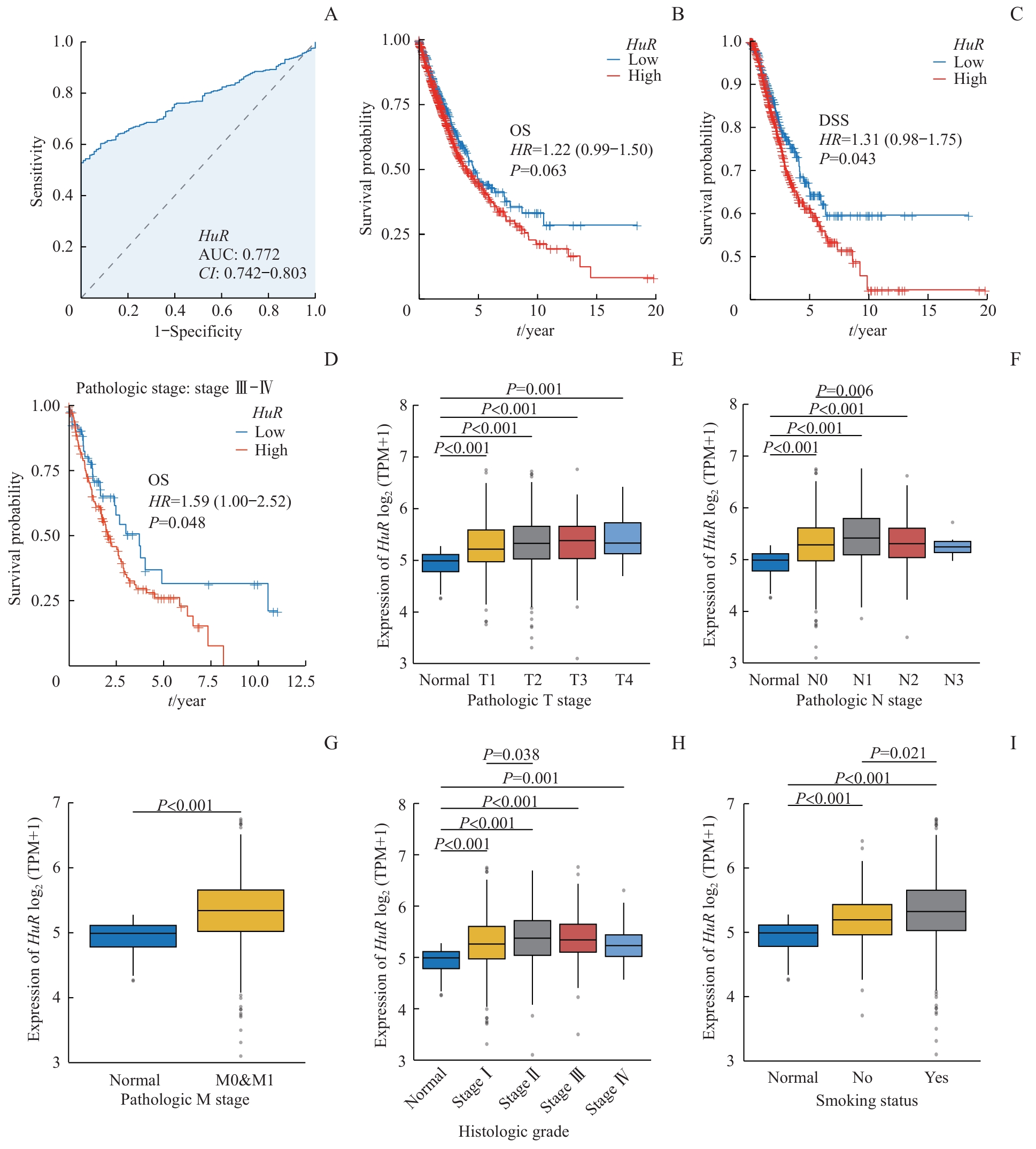
图2 HuR 表达对肺癌的诊断与预后价值Note: A. ROC curve showing the diagnostic potential of HuR in lung cancer. B‒D. Kaplan-Meier survival analyses comparing high and low HuR expression groups: overall survival (OS) in all patients (B); DSS in all patients (C); OS in stage Ⅲ‒Ⅳ patients (D). E‒I. Associations between HuR expression and T stage (E), N stage (F), M stage (G), histologic grade (H), and smoking status (I) in lung cancer patients.
Fig 2 Diagnostic and prognostic value of HuR expression in lung cancer
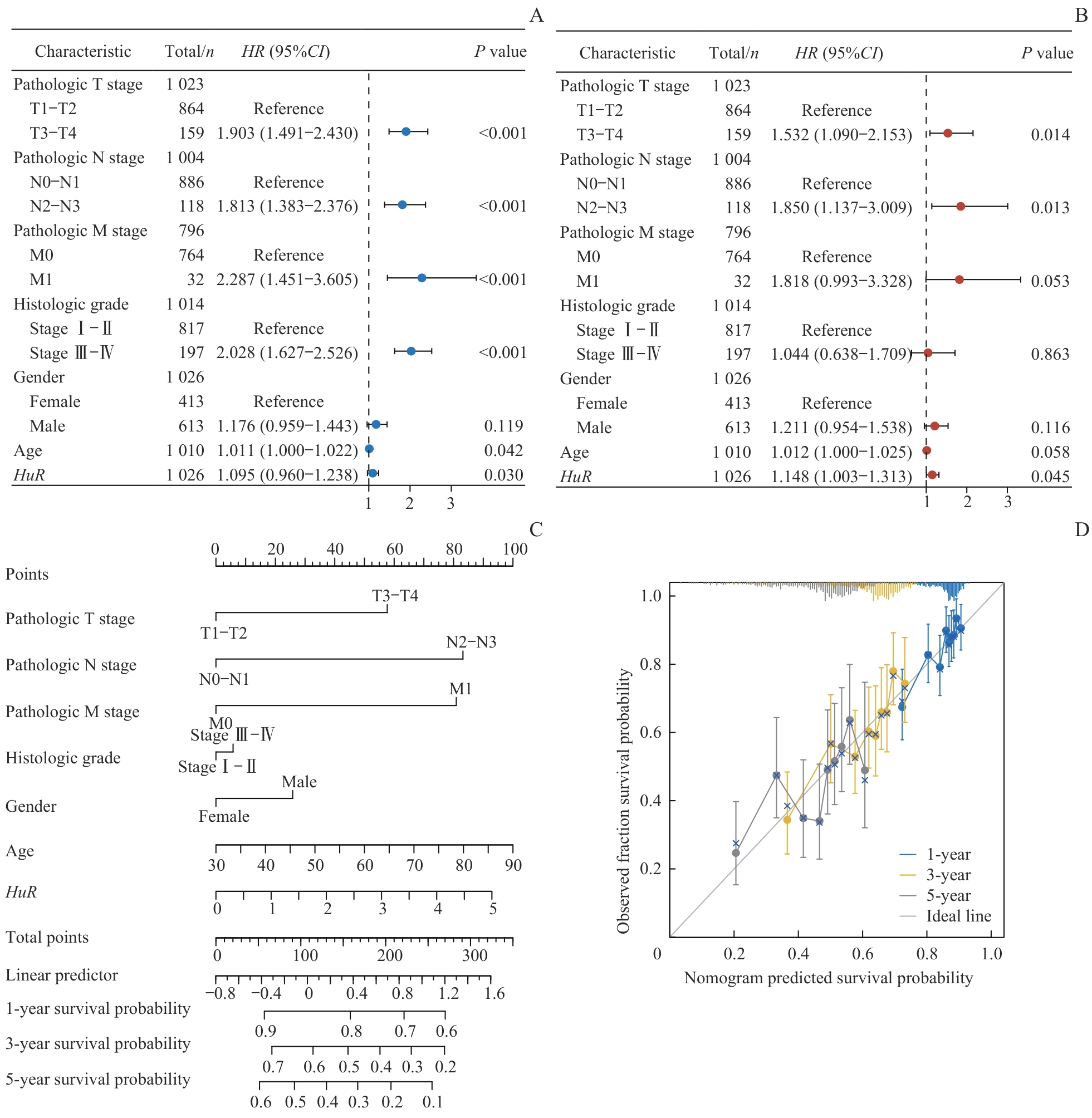
图3 肺癌预后影响因素分析及预后模型构建Note: A/B. Forest plots of univariate (A) and multivariate (B) Cox regression analyses of clinicopathological characteristics in lung cancer patients. C. Prognostic nomogram integrating clinical variables and HuR expression for predicting 1-, 3-, and 5-year OS in lung cancer patients. D. Calibration curves of the prognostic model.
Fig 3 Analysis of prognostic factors and construction of a prognostic model in lung cancer
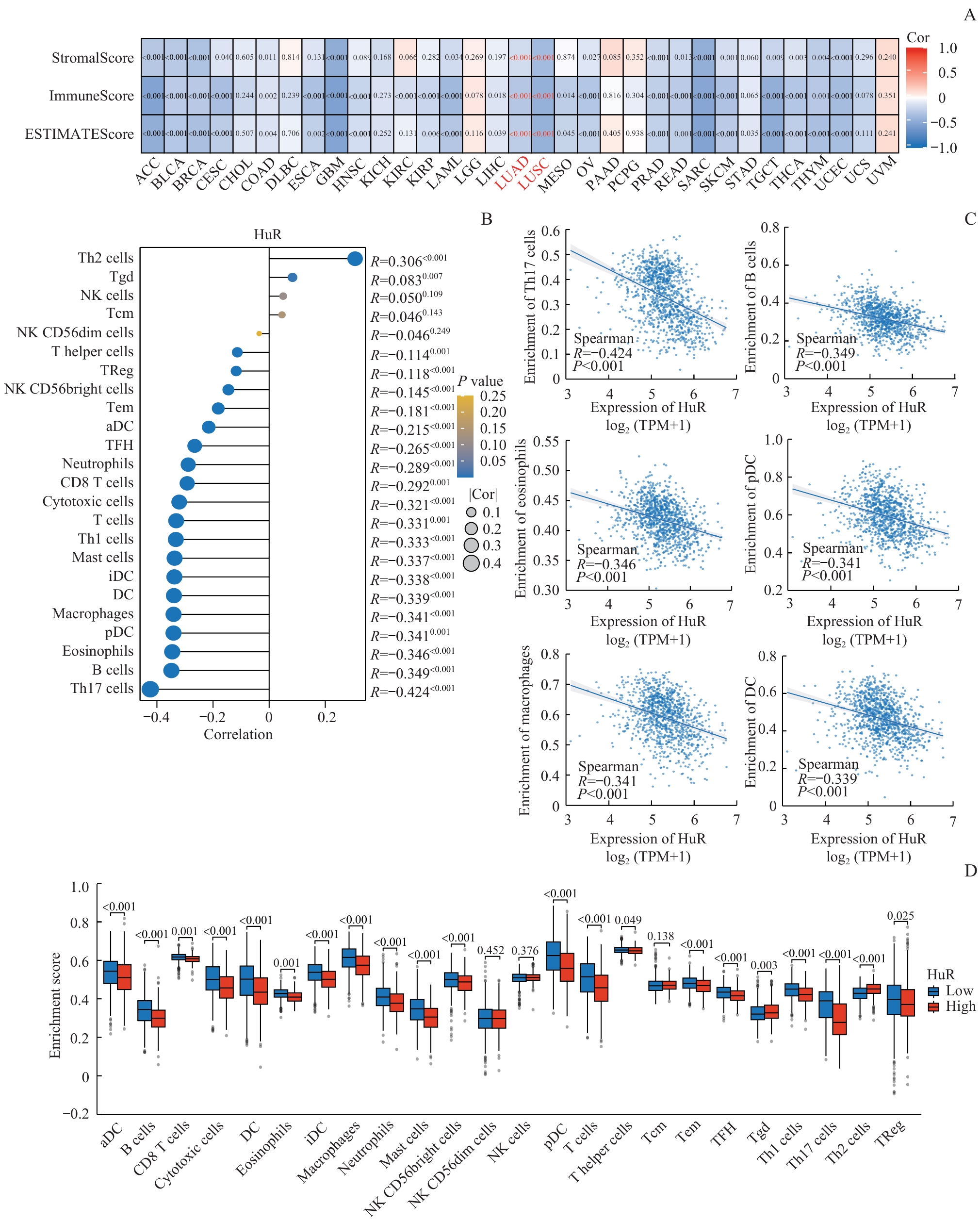
图4 HuR表达与肿瘤免疫微环境的负相关性Note: A. Heatmaps showing correlations between HuR expression and ImmuneScore, StromalScore, and ESTIMATEScore across 33 cancer types. B. Lollipop plot depicting the correlation between HuR expression and the infiltration levels of 24 immune cell types in lung cancer. C. Scatter plots illustrating the correlations between HuR expression and specific immune cell infiltration levels. D. Comparison of immune infiltration scores for 24 immune cell types between HuR high- and low-expression groups in lung cancer. Cor—correlation. NK cells—natural killer cells; Tgd—γδT cells; Tcm—central memory T cells; TReg—regulatory T cells; Tem—effector memory T cells; TFH—follicular helper T cells; DC—dendritic cells; aDC—activated dendritic cells; iDC—immature dendritic cells; pDC—plasmacytoid dendritic cells.
Fig 4 Negative correlation between HuR expression and the tumor immune microenvironment in lung cancer
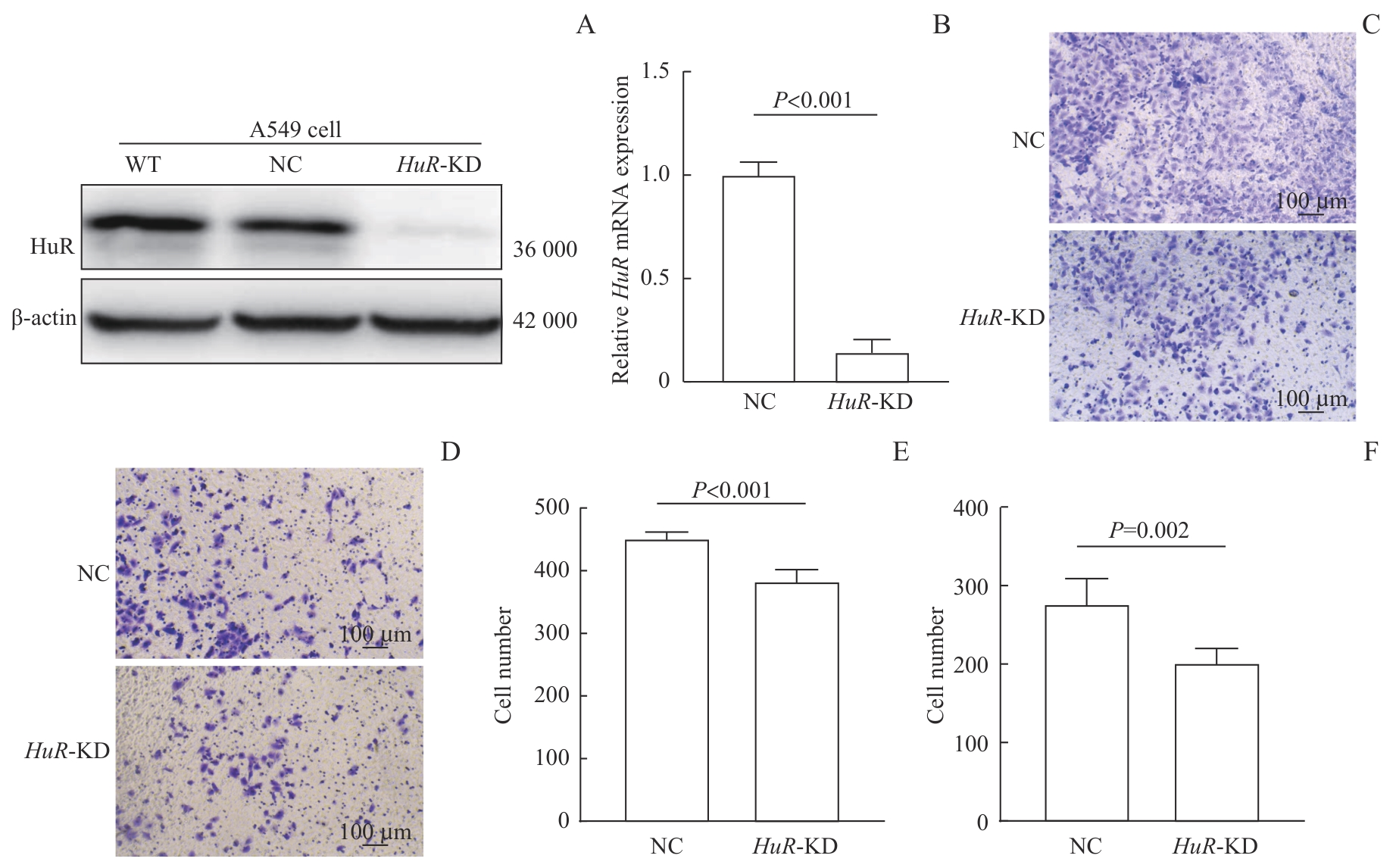
图5 HuR 敲低对A549细胞迁移与侵袭的抑制效果Note: A. Western blotting analysis of A549 wild-type (WT) cells, negative-control (NC) cells, and A549 HuR-KD cells. B. Relative mRNA expression of HuR in Lipofectamine 2000-treated NC cells and A549 HuR-KD cells. C‒F. Representative images (C, D) and quantitative analyses (E, F) of migration (C, E) and invasion (D, F) assays in A549 HuR-KD vs NC cells.
Fig 5 Inhibitory effect of HuR knockdown on the migration and invasion of A549 cells
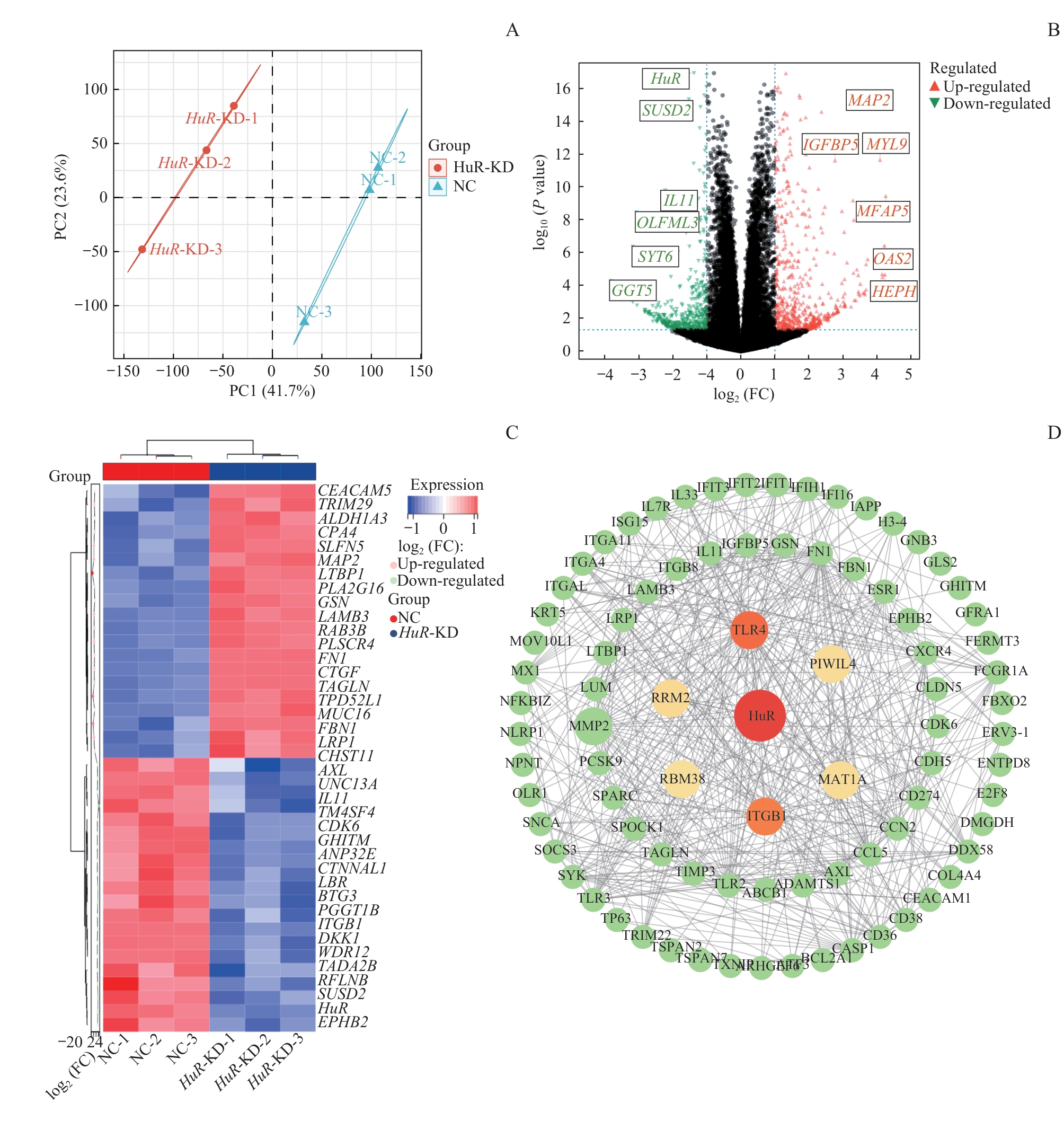
图6 HuR 敲低诱导的转录组重编程与 ITGB1 下调Note: A. PCA plot of RNA-seq samples from HuR-KD and A549 WT cells. B/C. Volcano plot (B) and heatmap (C) of DEGs between HuR-KD and A549 WT cells. D. PPI network of identified DEGs. Nodes represent DEGs; edges indicate functional interactions between nodes. FC—fold change.
Fig 6 Transcriptional reprogramming and ITGB1 downregulation induced by HuR knockdown
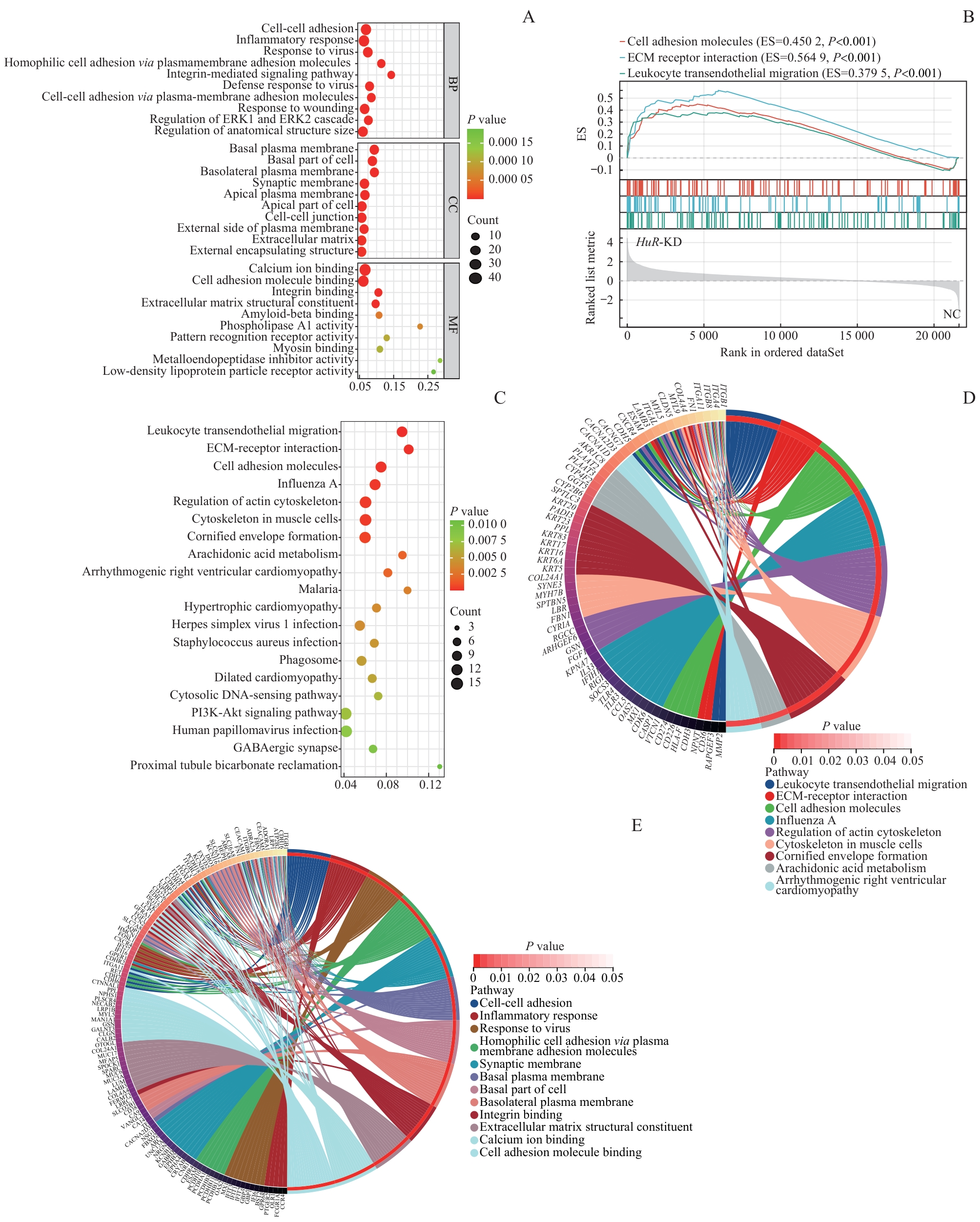
图7 功能富集分析揭示HuR通过ITGB1调控肺癌微环境的潜在机制Note: A. Bubble plot of GO enrichment analysis of DEGs after HuR-KD in lung cancer cells. B. Chord plot visualizing GSEA results. ES—Enrichment score. C. Bubble plot of KEGG pathway enrichment analysis. D. Circle plot displaying KEGG functional enrichment. E. Circle plot displaying GO functional enrichment.
Fig 7 Functional enrichment analysis reveals the potential mechanism of HuR in regulating the lung cancer microenvironment via ITGB1
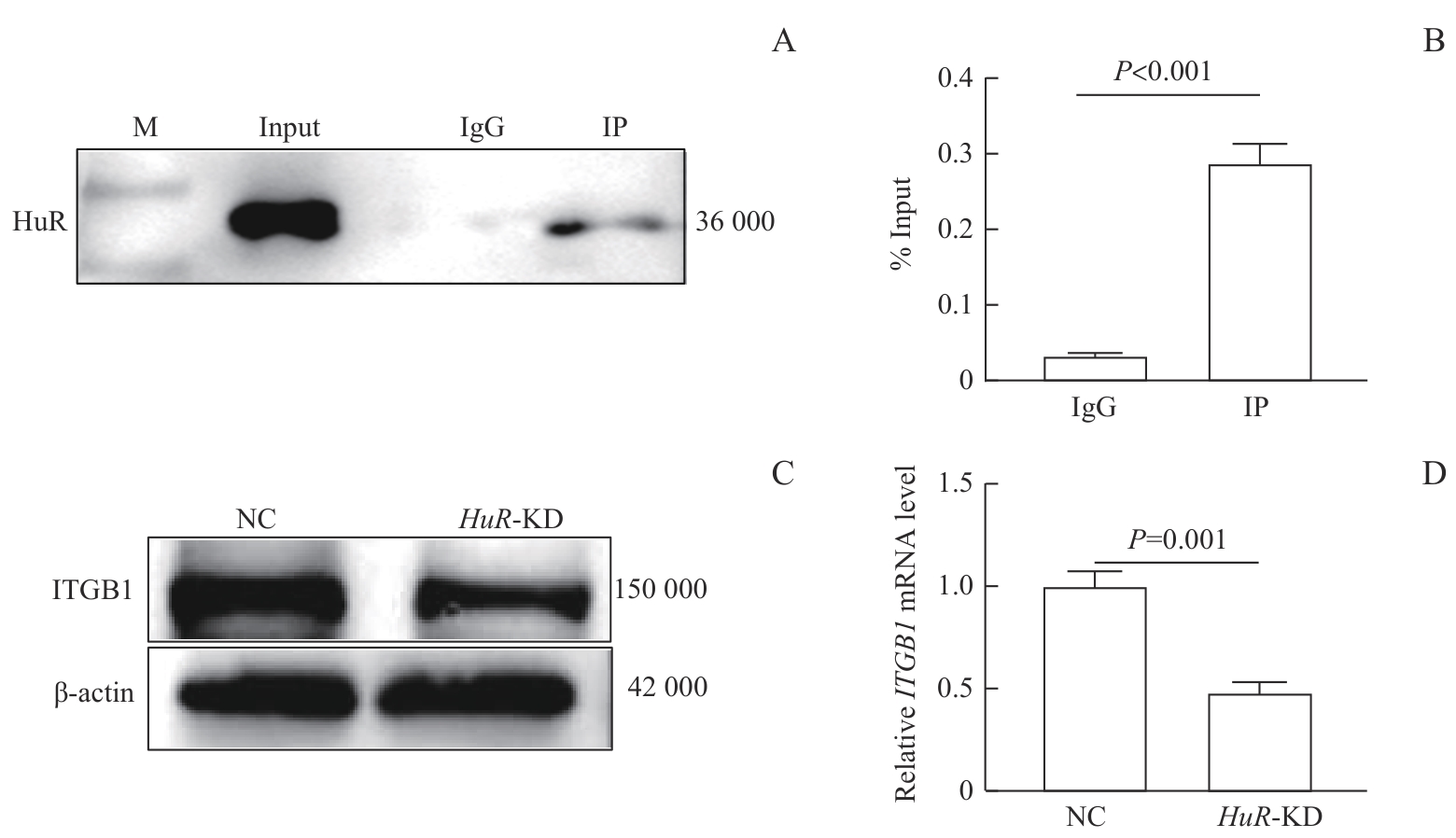
图8 HuR与ITGB1的分子相互作用及表达调控验证Note: A. Western blotting analysis of target protein HuR after RIP, showing the expression levels in the Input, IgG control, and IP (anti-HuR antibody) groups. B. RIP-qPCR analysis of the binding between HuR protein and ITGB1 mRNA in A549 cells. RNA-protein complexes were immunoprecipitated using an anti-HuR antibody, and the enrichment of ITGB1 was detected by qPCR. C/D. Western blotting (C) and RT-qPCR (D) analyses showing that knockdown of HuR led to a decrease in both mRNA and protein levels of ITGB1 in A549 cells.
Fig 8 Validation of the molecular interaction and expression regulation between HuR and ITGB1
| [1] | Kuang Z Y, Wang J X, Liu K X, et al. Global, regional, and national burden of tracheal, bronchus, and lung cancer and its risk factors from 1990 to 2021: findings from the global burden of disease study 2021[J]. EClinicalMedicine, 2024, 75: 102804. |
| [2] | Sung H, Ferlay J, Siegel R L, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin, 2021, 71(3): 209-249. |
| [3] | Wu X Q, Xu L. The RNA-binding protein HuR in human cancer: a friend or foe [J]. Adv Drug Deliv Rev, 2022, 184: 114179. |
| [4] | Sidali A, Teotia V, Solaiman N S, et al. AU-rich element RNA binding proteins: at the crossroads of post-transcriptional regulation and genome integrity[J]. Int J Mol Sci, 2021, 23(1): 96. |
| [5] | Clark M E, Farinha A, Morrison A R, et al. Structural, biological, and biomedical implications of mRNA interactions with the master regulator HuR[J]. NAR Mol Med, 2025, 2(1): ugaf002. |
| [6] | Yadav V, Singh T, Sharma D, et al. Unraveling the regulatory role of HuR/microRNA axis in colorectal cancer tumorigenesis[J]. Cancers, 2024, 16(18): 3183. |
| [7] | Kassabri L, Benhamou R I. Druglike molecular degraders of the oncogenic RNA-binding protein HuR[J]. JACS Au, 2025, 5(8): 3879-3891. |
| [8] | Raguraman R, Shanmugarama S, Mehta M, et al. Drug delivery approaches for HuR-targeted therapy for lung cancer[J]. Adv Drug Deliv Rev, 2022, 180: 114068. |
| [9] | Shatat A S, Mahgoup E M, Rashed M H, et al. Molecular mechanisms of extracellular-ATP-mediated colorectal cancer progression: implication of purinergic receptors-mediated nucleocytoplasmic shuttling of HuR[J]. Purinergic Signal, 2024, 20(6): 669-680. |
| [10] | Finan J M, Guo Y F, Bartlett A Q, et al. HuR-regulated extracellular vesicles promote endothelial cell remodeling in pancreatic cancer[J]. Cancer Res Commun, 2025, 5(9): 1501-1515. |
| [11] | Peng J, Quan J C, Wang X R. Integrated pan-cancer analysis of RNA binding protein HuR investigates its biomarker potential in prognosis, immunotherapy, and drug sensitivity[J]. PLoS Comput Biol, 2025, 21(8): e1013374. |
| [12] | Meng L J, Wu H T, Wu J X, et al. Aberrant CircTMEM45A facilitates inflammatory progression of esophageal squamous cell carcinoma through m5C-mediated NLRP3 activation[J]. Cancer Res, 2025, 85(14): 2694-2713. |
| [13] | Sun L, Guo S W, Xie Y P, et al. The characteristics and the multiple functions of integrin β1 in human cancers[J]. J Transl Med, 2023, 21(1): 787. |
| [14] | Huang Q L, Wang J, Ning H J, et al. Integrin β1 in breast cancer: mechanisms of progression and therapy[J]. Breast Cancer, 2025, 32(1): 43-59. |
| [15] | Lee J H, Son S, Ko Y, et al. Nidogen-1 suppresses cell proliferation, migration, and glycolysis via integrin β1-mediated HIF-1α downregulation in triple-negative breast cancer[J]. Sci Rep, 2025, 15(1): 10633. |
| [16] | Ren R M, Zhang S, Peng Z, et al. Matrix stiffness regulates glucose-6-phosphate dehydrogenase expression to mediate sorafenib resistance in hepatocellular carcinoma through the ITGB1-PI3K/AKT pathway[J]. Cell Death Dis, 2025, 16(1): 538. |
| [17] | Su C, Mo J, Dong S L, et al. Integrinβ-1 in disorders and cancers: molecular mechanisms and therapeutic targets[J]. Cell Commun Signal, 2024, 22(1): 71. |
| [18] | Ye X, Fu Q, Xiao H. The role of RNA-binding protein HuR in lung cancer by RNA sequencing analysis[J]. Front Genet, 2022, 13: 813268. |
| [19] | Zhou X, Han J S, Zuo A N, et al. THBS2+ cancer-associated fibroblasts promote EMT leading to oxaliplatin resistance via COL8A1-mediated PI3K/AKT activation in colorectal cancer[J]. Mol Cancer, 2024, 23(1): 282. |
| [20] | Lu Y B, Chen Q Z, Zhu S, et al. Hypoxia promotes immune escape of pancreatic cancer cells by lncRNA NNT-AS1/METTL3-HuR-mediated ITGB1 m6A modification[J]. Exp Cell Res, 2023, 432(2): 113764. |
| [21] | Liu J X, Tang L, Chu W Z, et al. Cellular retinoic acid binding protein 2 (CRABP2), up-regulated by HPV E6/E7, leads to aberrant activation of the integrin β1/FAK/ERK signaling pathway and aggravates the malignant phenotypes of cervical cancer[J]. Biochem Genet, 2024, 62(4): 2686-2701. |
| [1] | 陈蓉, 张锰, 朱荻绮, 郭颖, 沈捷. 基于抗中性粒细胞胞质抗体的列线图模型对川崎病患儿并发冠状动脉病变风险的预测作用[J]. 上海交通大学学报(医学版), 2025, 45(4): 459-467. |
| [2] | 陈佳莹, 褚以忞, 彭海霞. 结直肠癌无进展生存时间预测模型及影响因素研究[J]. 上海交通大学学报(医学版), 2025, 45(3): 324-334. |
| [3] | 张先洲, 杜凤麟, 吴雷, 任逸喆, 赵明娜, 娄加陶. OGT通过ERK信号通路促进非小细胞肺癌增殖的机制研究[J]. 上海交通大学学报(医学版), 2025, 45(10): 1288-1297. |
| [4] | 朱鸣阳, 许元元, 任江浩, 黄嘉正, 李若楠, 谭强. 以磨玻璃结节为表现的肺腺癌亚肺叶切除研究综述[J]. 上海交通大学学报(医学版), 2024, 44(7): 922-927. |
| [5] | 刘晨茜, 韩林, 杨轶, 周韩, 刘亚云, 盛德乔. GPR87通过激活RHO/ROCK通路促进非小细胞肺癌的侵袭和迁移[J]. 上海交通大学学报(医学版), 2024, 44(12): 1514-1525. |
| [6] | 俞思薇, 徐梓淇, 陶梦玉, 范广建. 脂肪酸代谢紊乱通过上调ZNF143促进胰腺癌进展的机制研究[J]. 上海交通大学学报(医学版), 2024, 44(10): 1255-1265. |
| [7] | 杜少倩, 陶梦玉, 曹源, 王红霞, 胡孝渠, 范广建, 臧丽娟. CXCL9在乳腺癌中的表达及其与肿瘤免疫浸润特征的相关性研究[J]. 上海交通大学学报(医学版), 2023, 43(7): 860-872. |
| [8] | 黄华艳, 徐张闻笛, 夏立亮, 虞永峰, 陆舜. 表皮生长因子受体突变型晚期非小细胞肺癌免疫治疗的研究进展[J]. 上海交通大学学报(医学版), 2023, 43(5): 611-618. |
| [9] | 田晓梵, 董怡, 楼文晖, 张琪, 邱艺杰, 左丹, 王文平. 基于超声剪切波弹性成像参数与临床风险因素的术后胰瘘改良预测模型[J]. 上海交通大学学报(医学版), 2023, 43(4): 437-444. |
| [10] | 赵卓明, 刘振浩, 鲁曼曼, 张钰, 许林锋, 谢鹭. 基于TCR组库分析流程的非小细胞肺癌特征分析[J]. 上海交通大学学报(医学版), 2023, 43(12): 1520-1528. |
| [11] | 廖雅慧, 刘丽云, 朱泓睿, 林厚文, 严继舟, 孙凡. 海绵来源的smenospongine通过抑制非小细胞肺癌细胞中的EGFR-Akt-ABCG2信号通路抑制顺铂耐药[J]. 上海交通大学学报(医学版), 2022, 42(8): 997-1007. |
| [12] | 刘子杨, 王小文, 陈力. lncRNA GK-IT1通过调控醛缩酶A影响非小细胞肺癌细胞的恶性进展[J]. 上海交通大学学报(医学版), 2022, 42(5): 591-601. |
| [13] | 夏坤健, 邓林林, 王琳. 乳腺癌化学治疗致肝损伤预测模型的构建及其评价[J]. 上海交通大学学报(医学版), 2022, 42(4): 502-509. |
| [14] | 陆文清, 孟周文理, 虞永峰, 陆舜. 非小细胞肺癌第三代表皮生长因子受体-酪氨酸激酶抑制剂的耐药机制及治疗策略[J]. 上海交通大学学报(医学版), 2022, 42(4): 535-544. |
| [15] | 许静轩, 杜少倩, 曹源, 王红霞, 黄伟翼. MMP14在胰腺癌中的表达及其与肿瘤免疫微环境特征的相关性研究[J]. 上海交通大学学报(医学版), 2022, 42(3): 312-322. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||